All the tools you need to build better models, faster
Encord is the leading data development platform for advanced computer vision teams. Streamline your labeling and workflow management, intelligently clean and curate data, easily validate label quality, and evaluate model performance.
Trusted by pioneering AI teams
The complete data development platform for AI model development
Unlock the potential of AI for your organization. Streamline your data-centric AI pipelines to build better models and deploy high-quality production AI, faster.
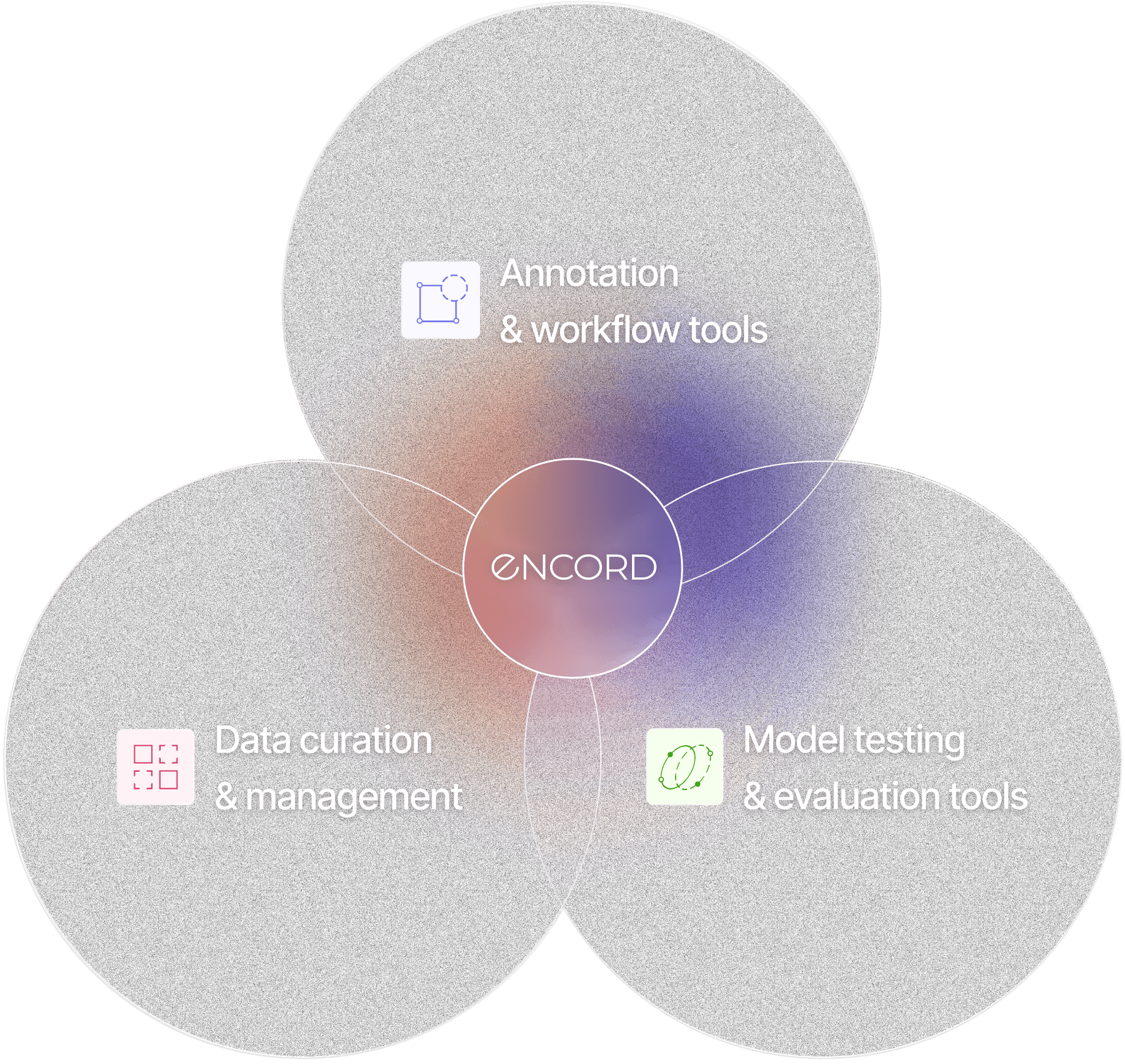
Encord AI Data Development Platform

Annotate
Annotation & workflow tools
Active
Model testing & evaluation

Index
Data curation & management
Understand and manage your visual data, prioritize data for labeling, and initiate active learning pipelines.
The complete data development platform for AI model development

Annotate
Scale and accelerate labeling workflows and teams.
annotation types
Supporting your labeling needs
Efficiently label any computer vision modality across image, video, medical imagery, or geospatial data and choose from a variety of tools to meet your annotation needs.

Image

Video

DICOM & NifTI

Synthetic-aperture radar

Documents
One-click automated labels
Leverage state-of-the-art foundational models to accelerate labeling projects and build high-quality training data faster.
Explore automated labeling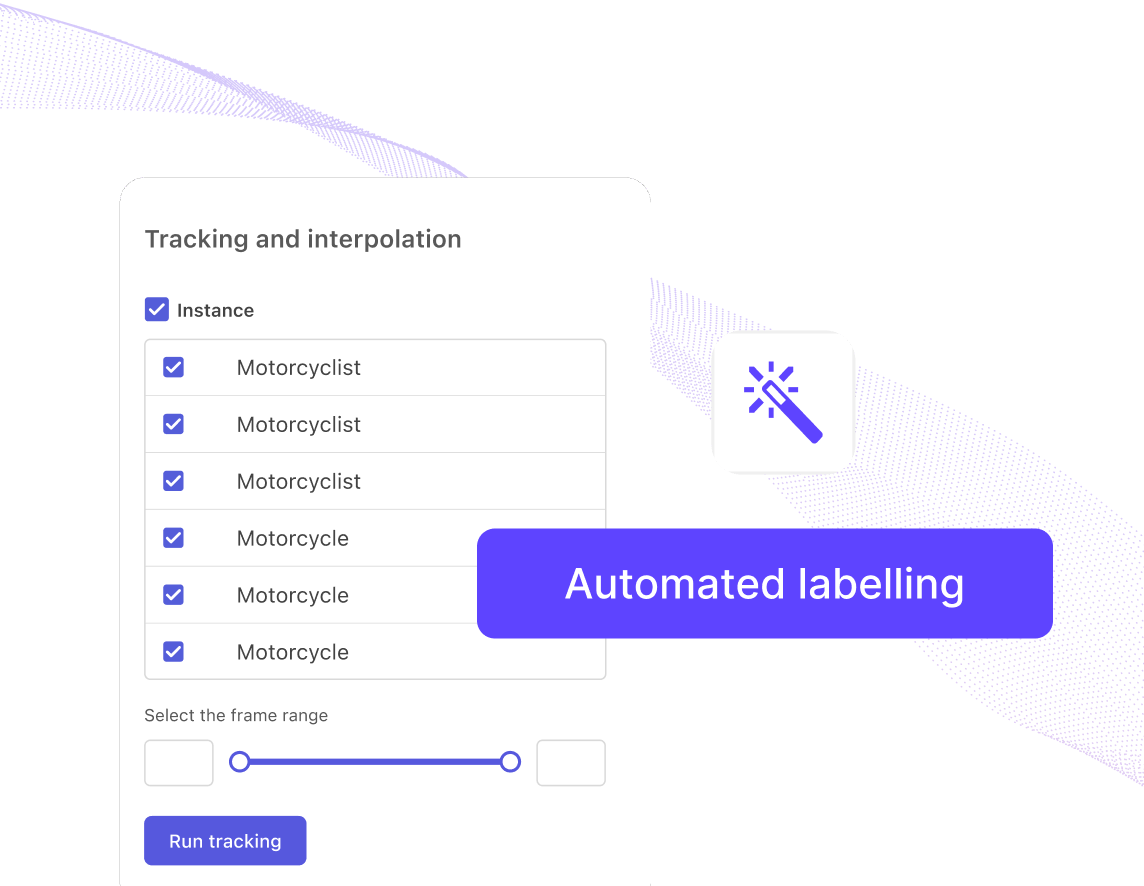
Customizable workflows, automated pipelines
Integrate humans-in-the-loop seamlessly to accelerate training data creating with fully customizable Workflows and expert review.
Explore workflows
One-click automated labels
Customizable workflows, automated pipelines

Streamlining data operations
Monitor team and annotator performance with actionable dashboards to ensure training data quality.

Increase labelling efficiency
Integrate active learning workflows to identify and label the data that boosts model performance.

On-demand expert labeling services
Access thousands of expert labelers to scale your projects without compromising data quality.

Active
Monitor, troubleshoot, and evaluate your models.
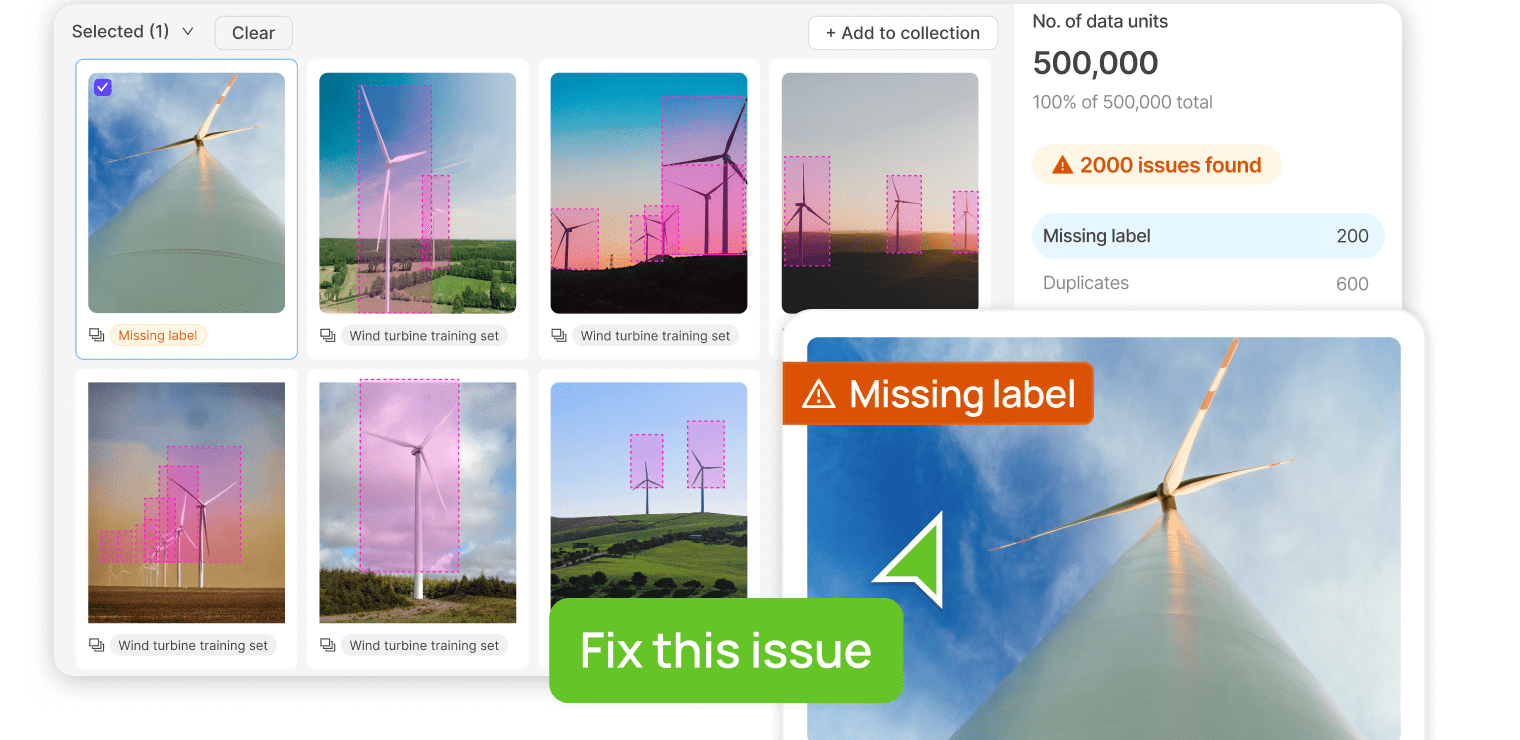
Find & fix data issues with ease
Build robust models with powerful error analysis

Fine-tune foundational models
Fine-tune foundation models to automate annotation on just a handful of labels without compromising on quality.

Evaluate model performance
Deconstruct model performance with automatic reporting on metrics like mAP, mAR, and F1 Score.

Understand model blind spots
Effortlessly uncover model edge-cases, underperforming clusters, underrepresented classes and more.


Index
Understand and manage your data.

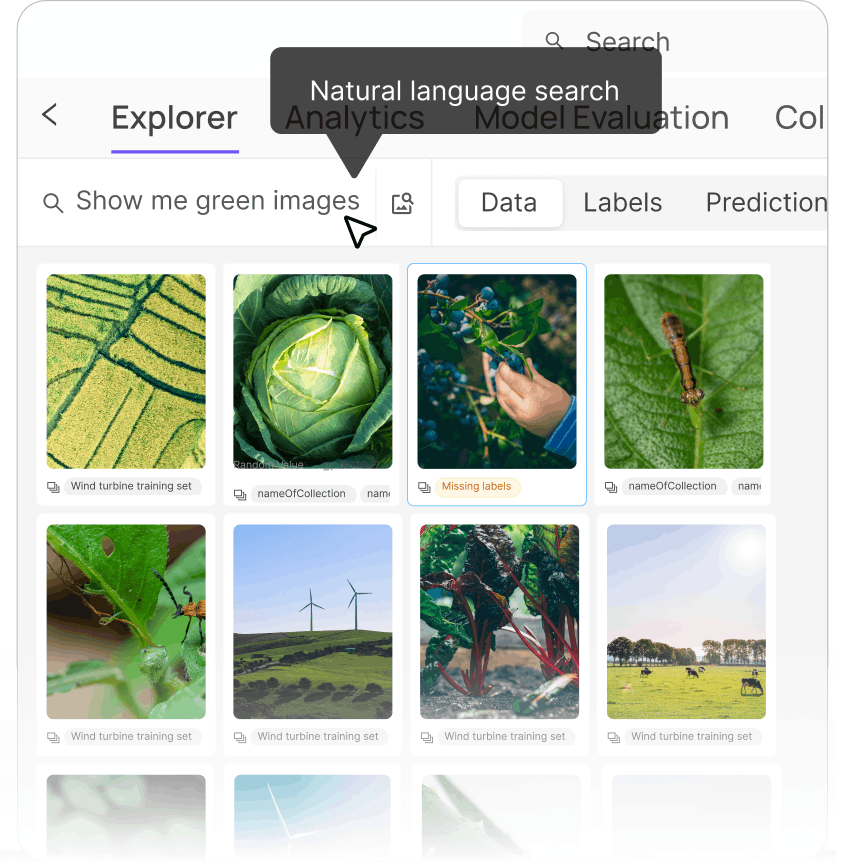
Explore your data with ease

Boost model performance with the right data

Support for all
key visual formats
Monitor team and annotator performance with actionable dashboards to ensure training data quality.

Integrate your data pipelines,
hassle free
Integrate active learning workflows to identify and label the data that boosts model performance.

Advanced filtering options
Filter data by brightness, RGB, contrast and many more visual quality metrics to curate more effective datasets to power your models.
testimonials
What our clients say

Integrations
Integrate seamlessly with your toolstack
Connect your secure cloud storage, MLOps tools, and much more with dedicated integrations that slot seamlessly into your workflows.

Security
Built with security in mind
Encord is SOC2, HIPAA, and GDPR compliant with robust security and encryption standards.

API/sdk
Developer-friendly for easy access
Leverage our API/SDK to programatically access projects, datasets & labels within the platform via API.
Take control of your ML pipeline with Encord
Forget fragmented workflows, annotation tools, and Notebooks for building AI applications. Encord's Data Development platform accelerates every step of taking your model into production.