Deploy medical AI faster with Encord
Level-up your model development and create high-quality training data up to 10x faster with the most advanced image labeling tool.









“Encord made it very easy to centrally keep track of annotations, including who had made them and who had reviewed them.”

Dr. James Ryan Mason
Neuroradiologist at Rapid AI
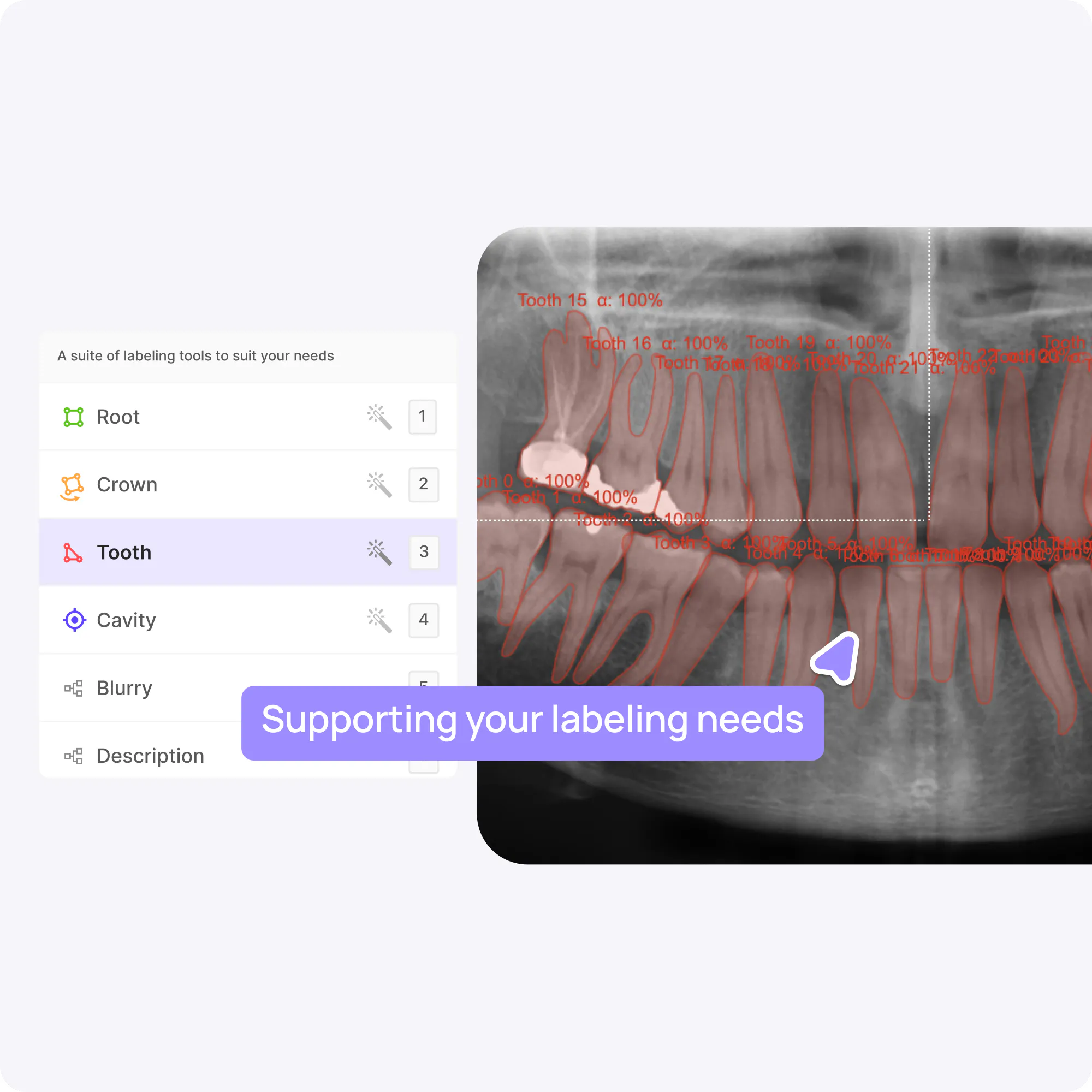
Pixel-perfect labeling at the speed of AI
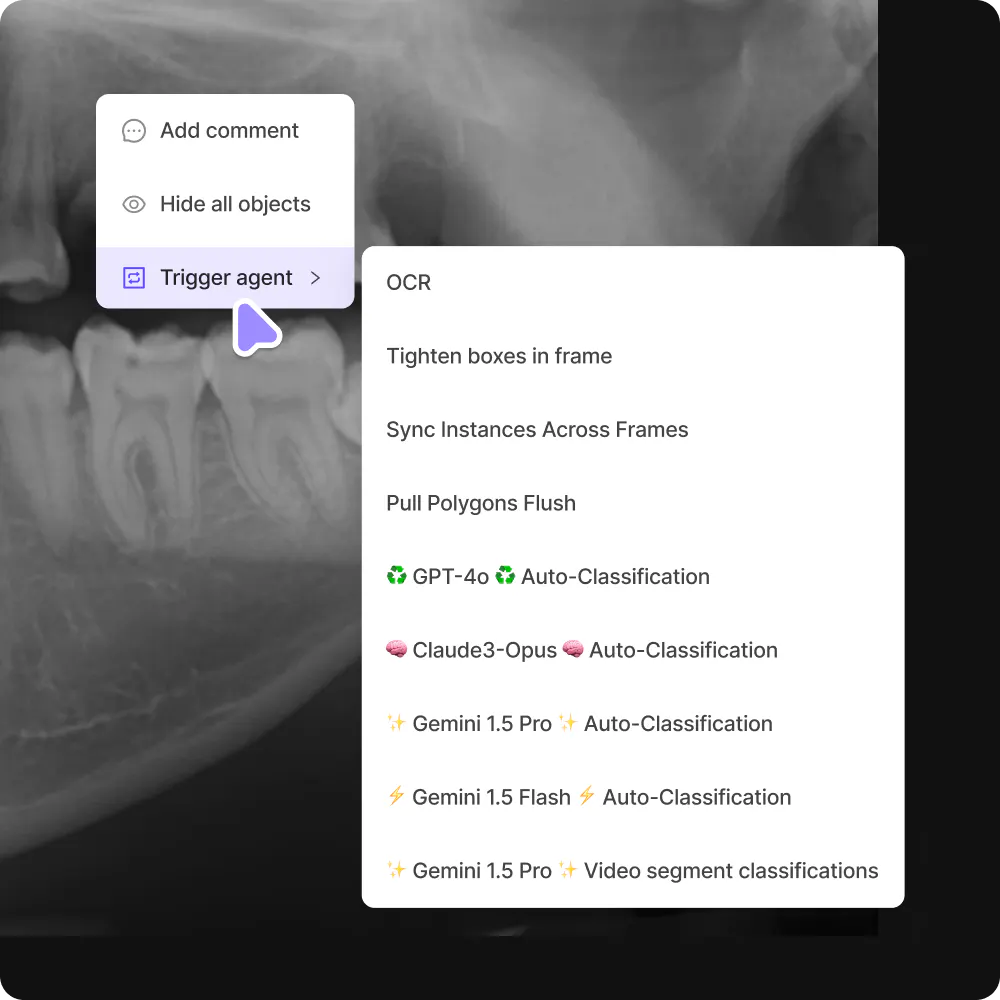
Experience an advanced, lightning-fast label editor with powerful automation capabilities to accelerate your labeling workflows.
A fully-customizable, medical-grade experience
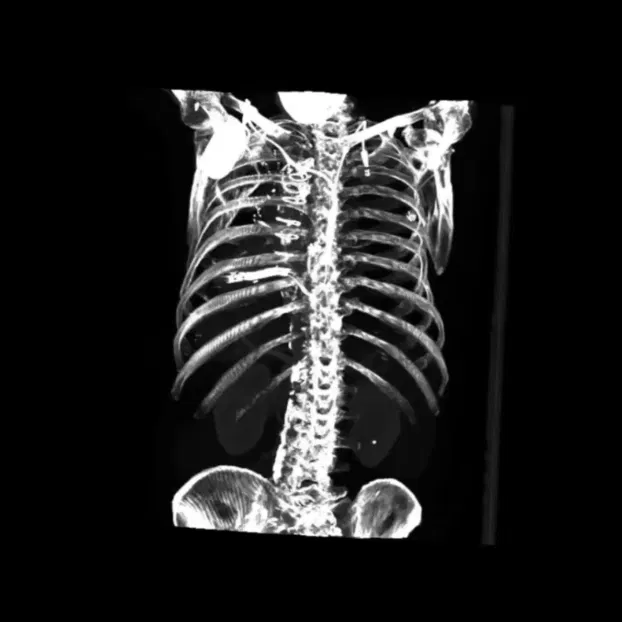
A truly 3D experience
Streamline your medical imaging workflows with 3D annotations for enhanced accuracy and multi-plane views for comprehensive annotation.
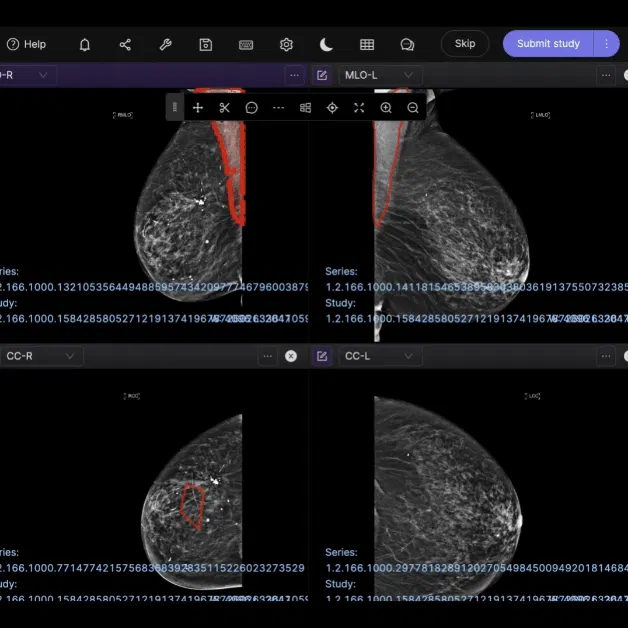
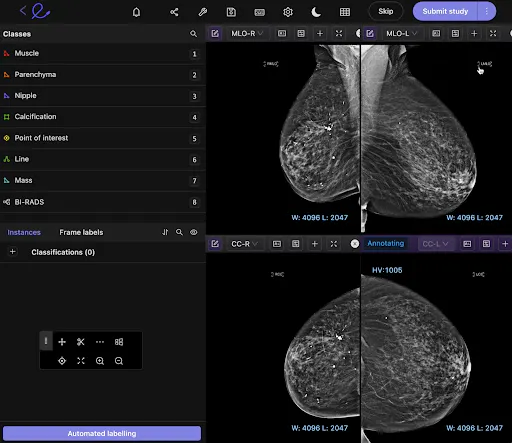
See it your way with hanging protocols
Customize your workspace to suit you with pre-built and customizable layouts to suit your annotation requirements.
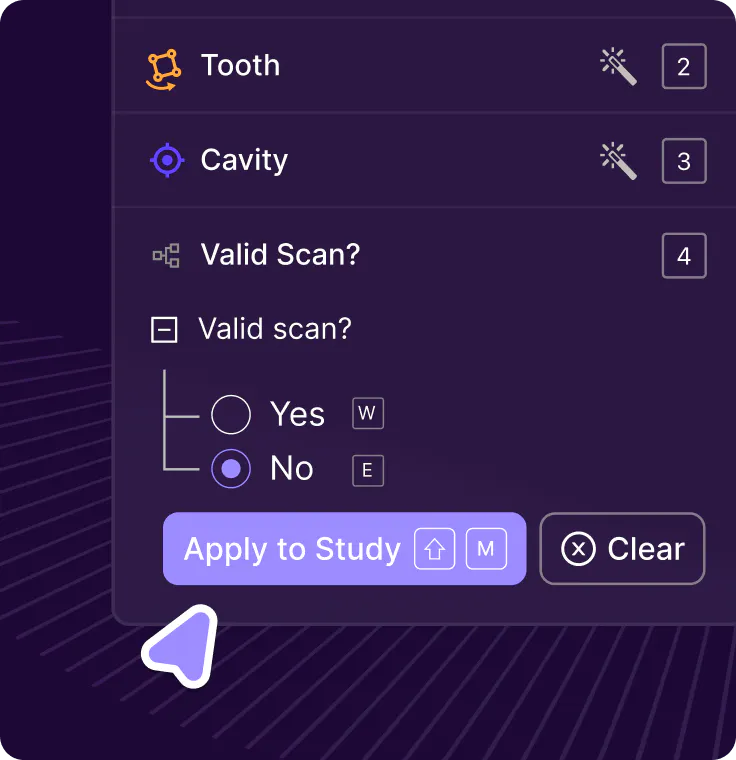
Study-level support & classifications
Encord automatically identifies series belonging to the same study using metadata and allows for classifications to be applied at the study-level.
Metadata viewer.Search, filter, and slice your data for all defined & custom metadata and sequences associated with your file.
Maximum intensity Projection.Simply drag and adjust the volume of your displayed images that suits your annotation needs.
HTJ2K Support.At the forefront of DICOM development, we support the latest transfer syntaxes for DICOM files.
Distance measurement.Measure real-world distances between any two points on an image to convert pixels into distance accurately.
Bring your custom models.Label data faster with models-in-the-loop. Bring your own model to Encord seamlessly.
Programmatically upload data.Use our API/SDK to programmatically upload your projects, datasets, and labels.
Testimonial
Trusted by leading AI teams
Customizable workflows, automated pipelines
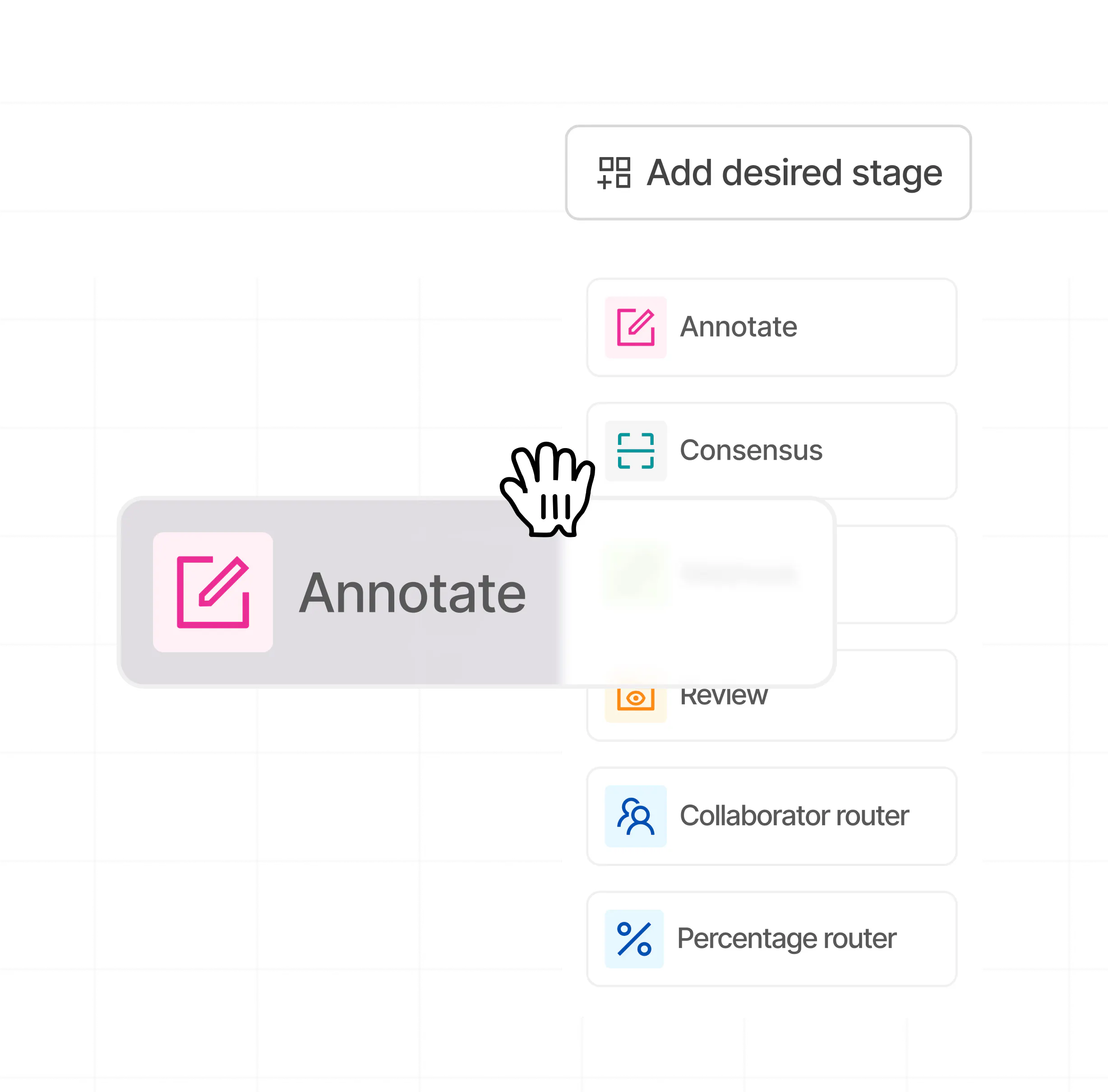
Build custom workflows with no-code
Add desired stages to your labeling workflow. Select from annotate, review, consensus, webhooks, and various routers to configure your workflows.
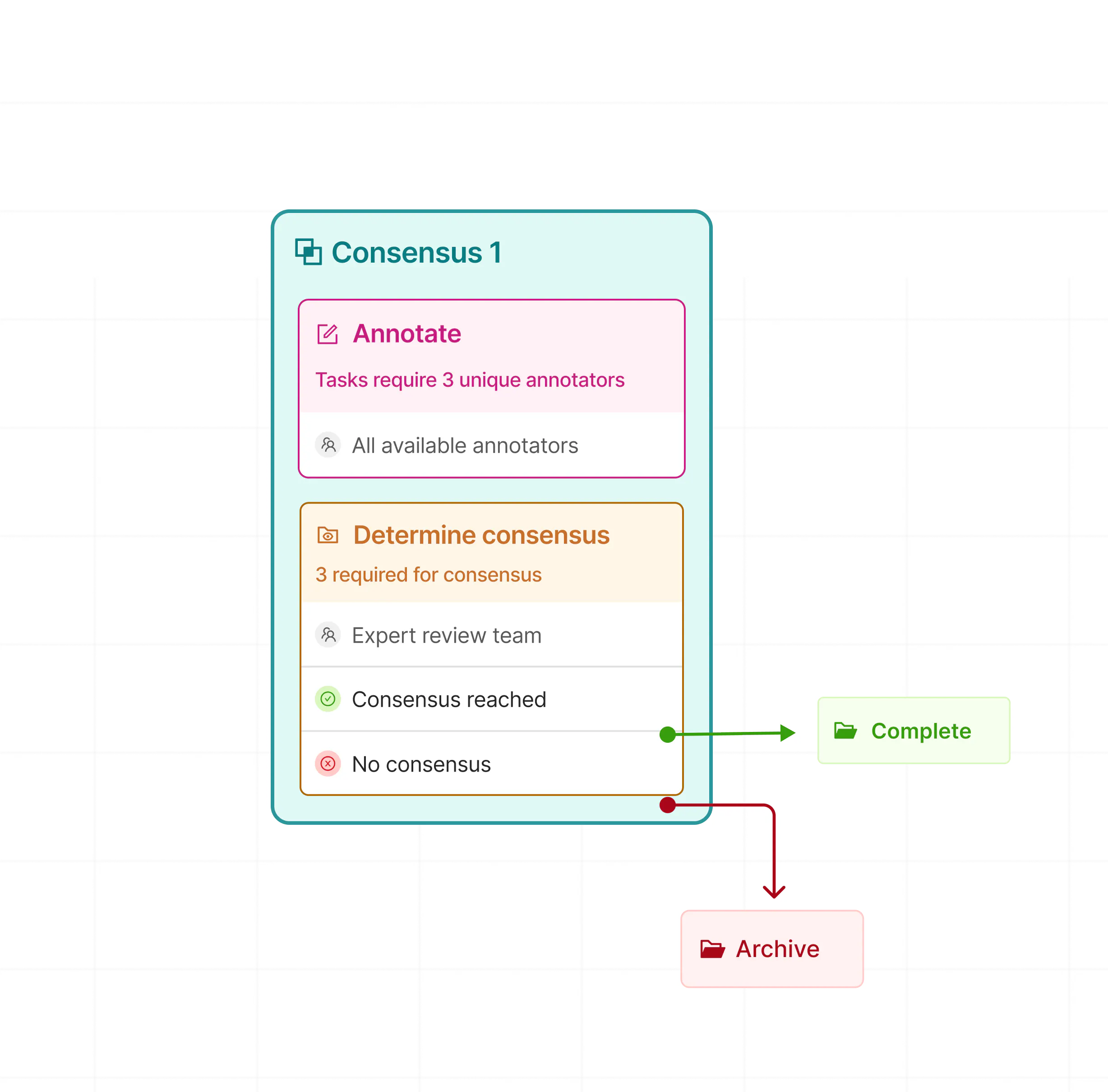
Ensure label accuracy with Consensus
Compare annotations between labellers and establish accepted thresholds that determine approved or rejected labels.
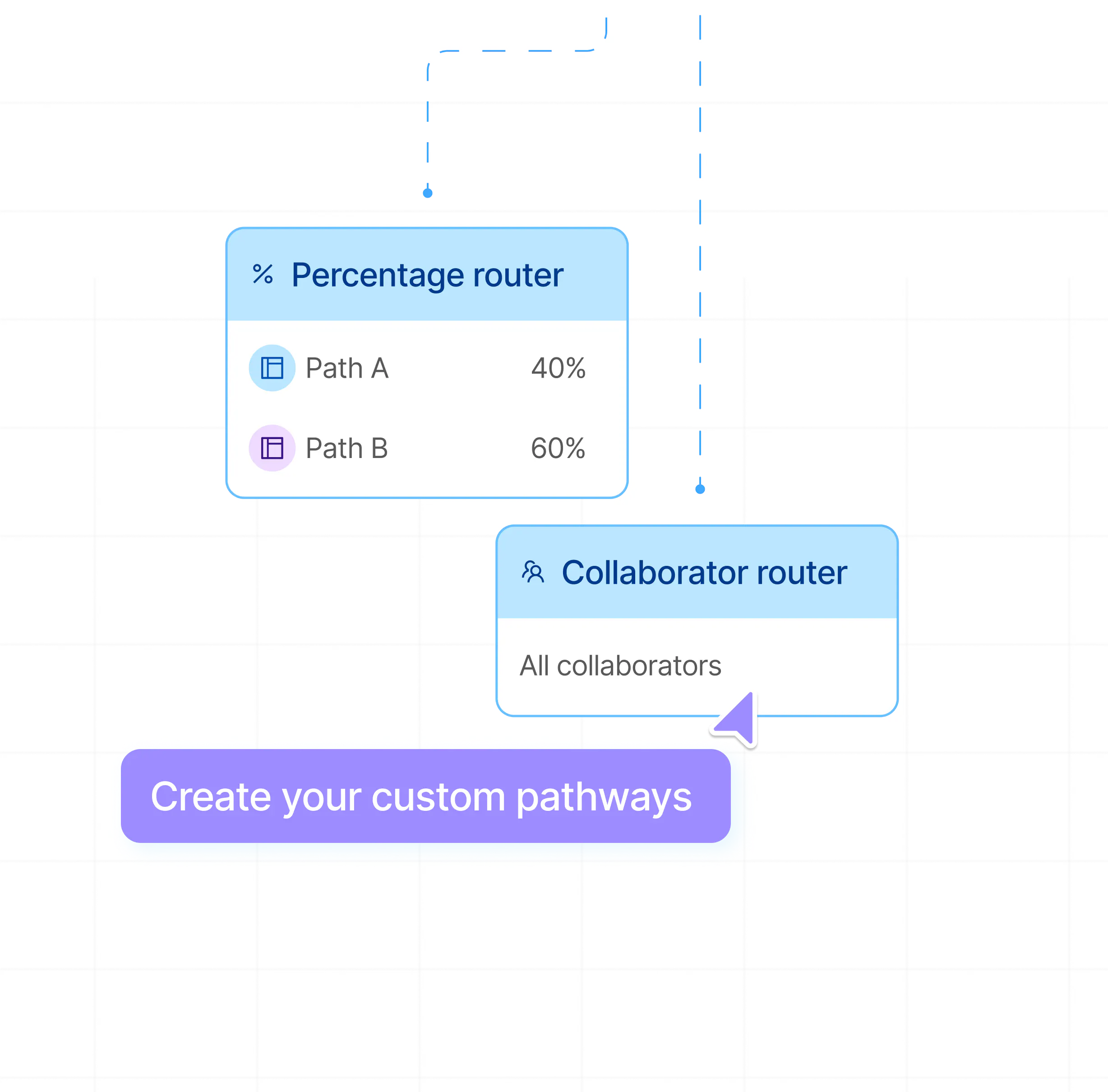
Custom pathways with advanced routers
Create specific pathways for any labeling instances. Leverage annotator-specific routing, label-specific routing, or weighted routing to suit your needs.
Built for scale
Effectively supporting datasets of 1,000 or 10,000,000 images
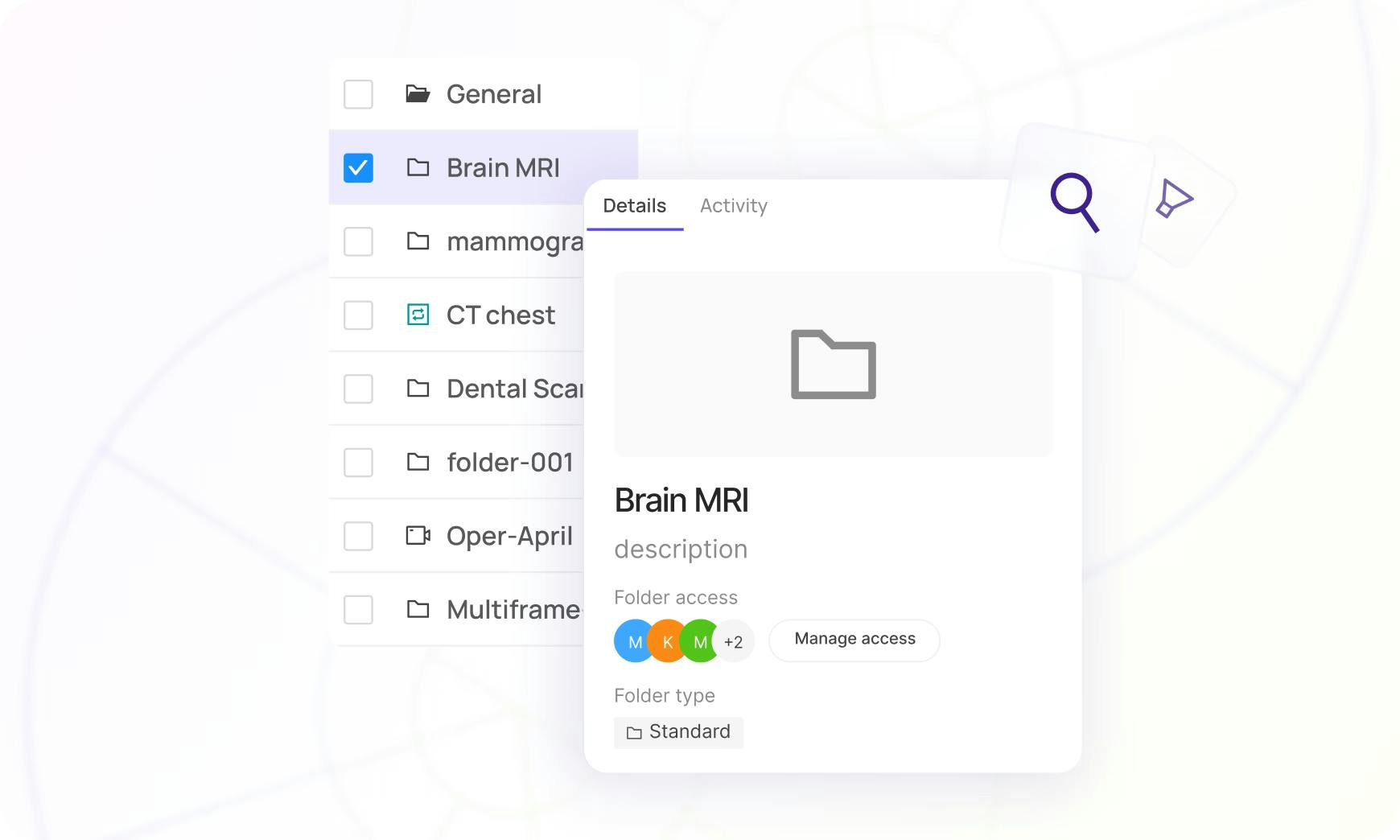
Build truly balanced datasets with ease
Filter and slice datasets in a consolidated visual explorer and export for labeling in one click. Encord supports deep search, filtering, and metadata analysis.

Build truly balanced datasets with ease
Filter and slice datasets in a consolidated visual explorer and export for labeling in one click. Encord supports deep search, filtering, and metadata analysis.
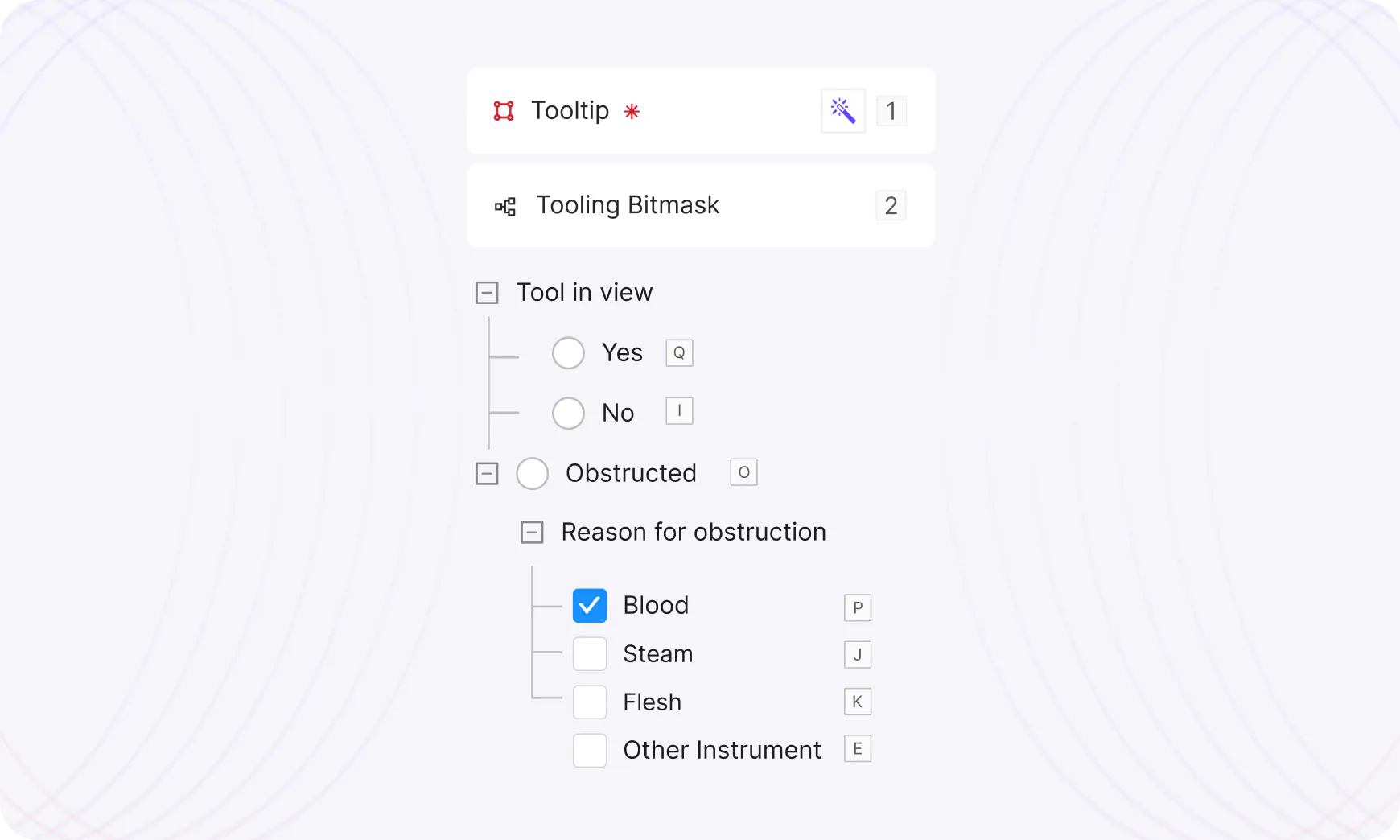
Rich knowledge structures enhance model context
Build nested relationship structures in your data schema to improve the quality of your model output.
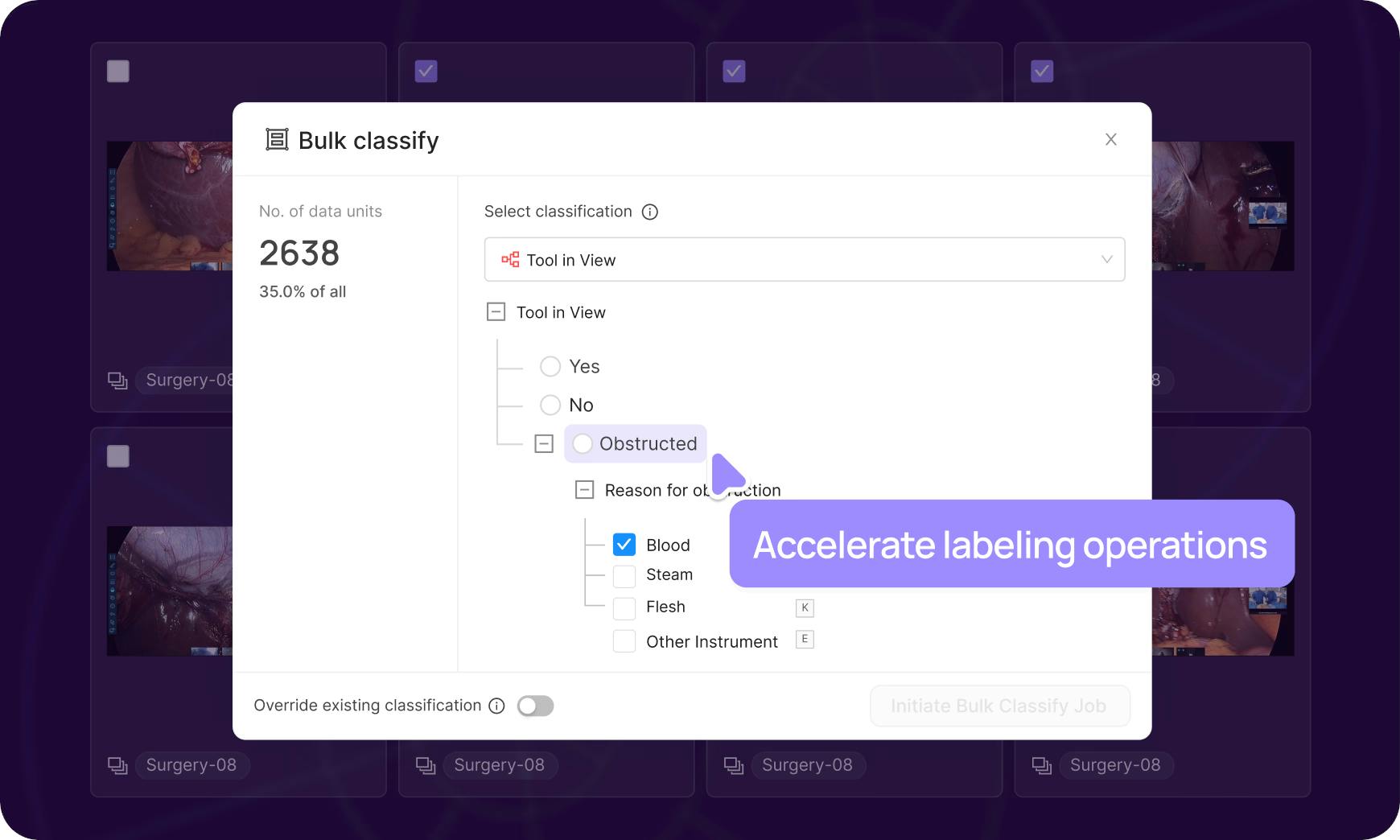
Bulk classifications
Leverage natural language or similarity search to select large datasets and label en masse, queue for review to drastically accelerate labeling operations.
Designed for your specific use case
A truly collaborative experience
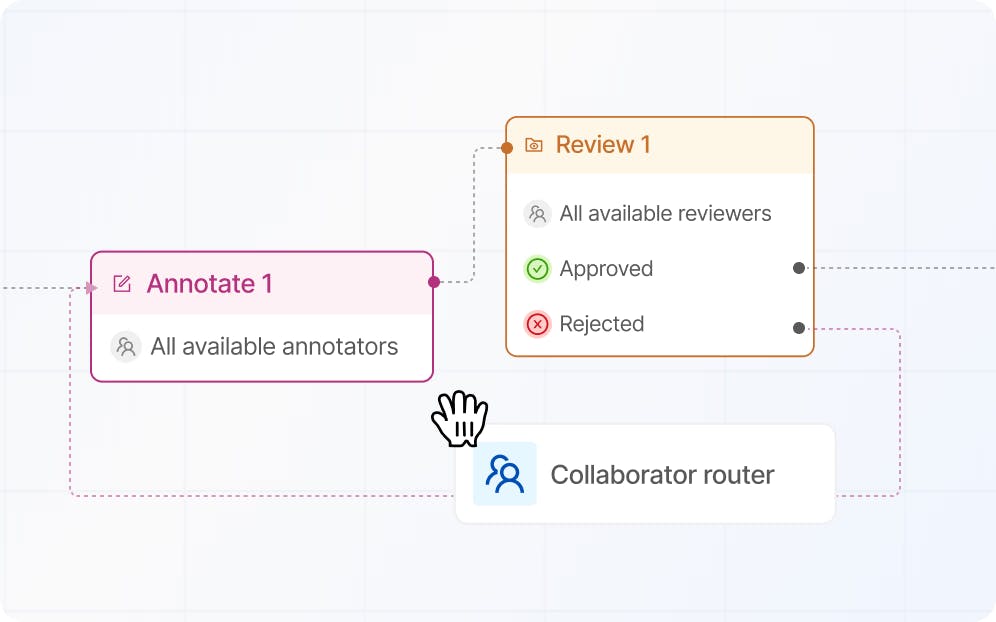
Build reliable quality control workflows
Build robust workflows with multi-step review stages, now with consensus.
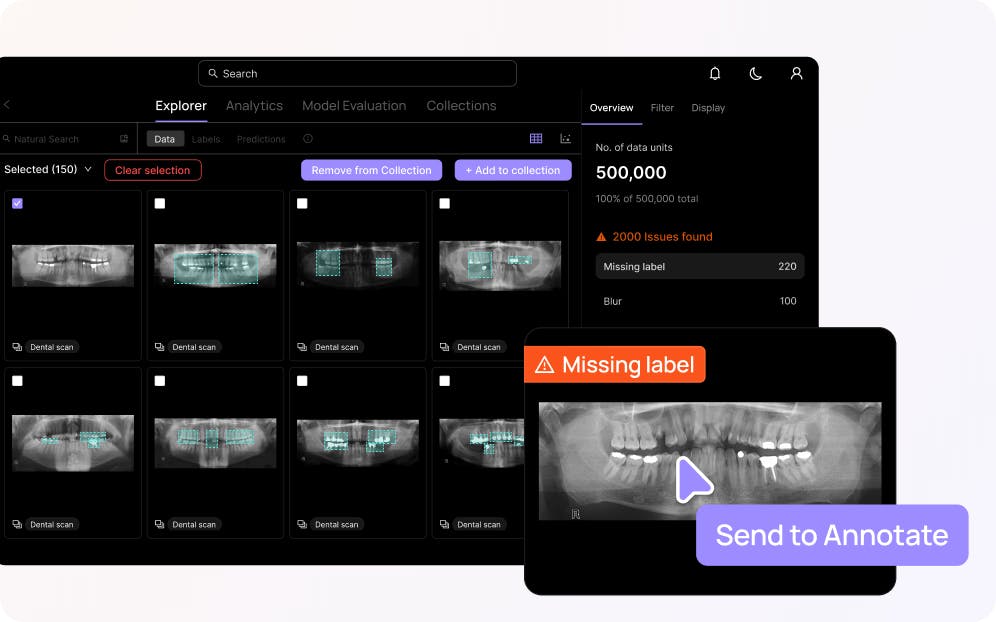
Find & fix label errors
Automatically surface labeling errors to shift your attention to the labels that are impacting model performance.
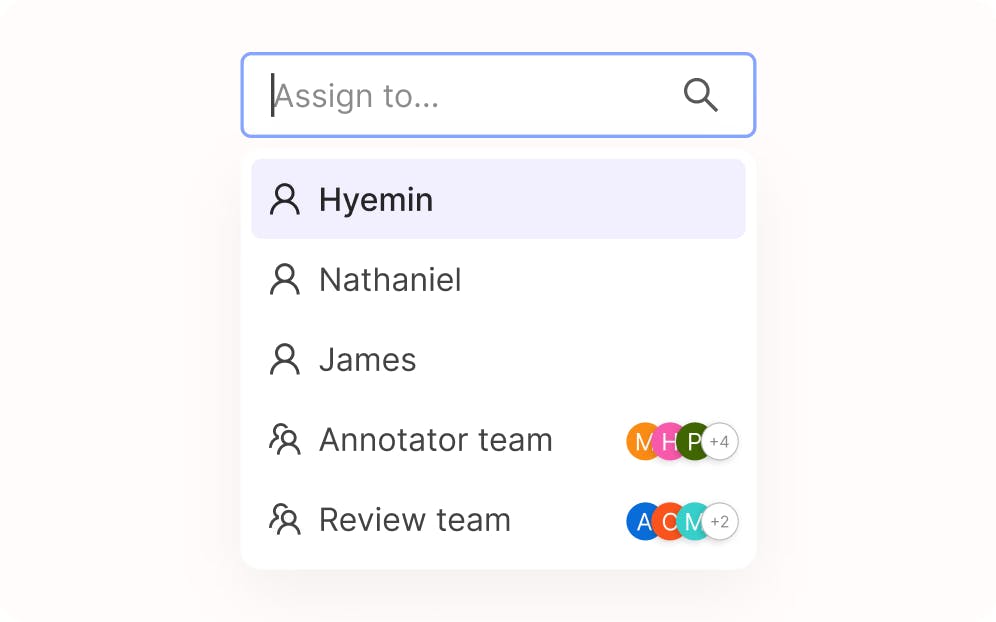
Collaborate with teams of all sizes
Seamlessly control user roles with permissioning, manage task assignment, and infinitely scale your MLOps workflows.

Collaborate with teams of all sizes
Seamlessly control user roles with permissioning, manage task assignment, and infinitely scale your MLOps workflows.
Enterprise-grade.
Built for scale.
Designed for reliable AI.
Built for scale.
Designed for reliable AI.
API/SDK-first. Zero data migration. Your data stays in your cloud.
Visit trust centre



Get the data right
300+ of the best AI teams in the world use Encord. Join them.