6 Best Open Source DICOM Annotation Tools
In this article, we will cover several of the most popular open-source tools for DICOM annotation 9 that our team often discusses with leaders from Data Operations and Machine Learning teams (as well as radiologists and clinicians), including the key use cases, benefits, and downsides of these tools – we will also look ahead to what’s next after getting started with these tools, and considerations that teams make as they go forward in their data annotation journey.
Viewing and annotating medical data, images, and videos is a crucial, and frequent, task for many practitioners in the healthcare industry. This is especially true as AI becomes more prominent across different fields of medicine. The AI in healthcare market is expected to grow from $11 billion in 2021 to $188 billion by 2030, which greatly increases the demand for high-quality medical data. (Source: Statista)
A starting point for many when evaluating how to go about this task, will be to start with open-source medical imaging annotation tools – these tools are a popular choice in the medical sector and can be a smart way to save money when getting started on an image or video dataset annotation project.
As we all know, there are pros and cons to using open-source tools for DICOM annotation projects. When conducting your own evaluation, it’s worth comparing what is on the market with what your own requirements are – based on your specific use cases and forward-looking plans.
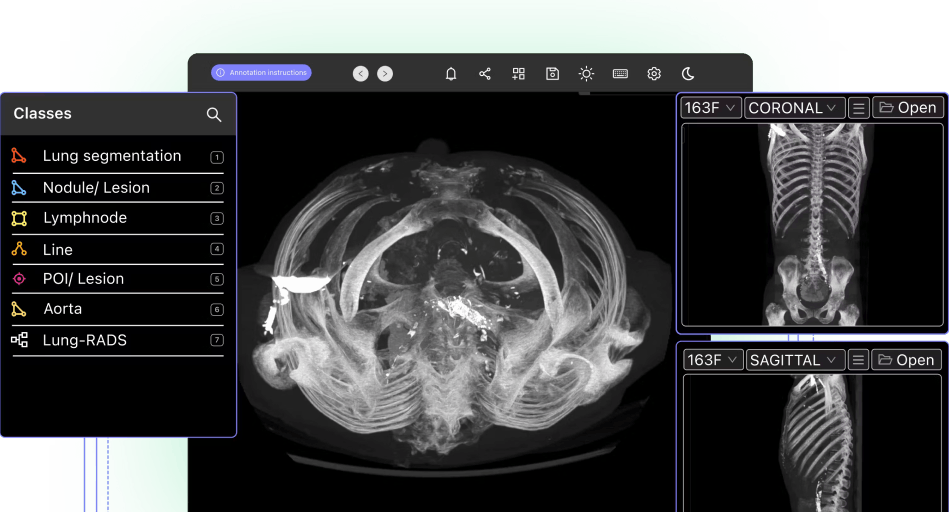
What are Open-Source Annotation Tools?
Open-source annotation tools are software programs whose source code is freely available for anyone to use. When we think of annotation platforms, what we mean by open-source annotation tools are tools that help teams with the broad annotation and labeling process (including use cases like image classification, image segmentation, data labeling, and object detection). They will be aimed at supporting almost any image or video annotation purpose (unless the license specifically prohibits a certain type of use).
Open-source tools are usually built collaboratively, with numerous — sometimes hundreds or thousands — of developers contributing to the source code. Tools are often tested using publicly available medical imaging datasets, and are usually financially supported by a charitable foundation, public/users donations, or one or more tech company sponsors.
What are The Advantages of Using Open-Source Annotation Tools for Medical Imaging?
The key advantages of open-source annotation tools for medical imaging are:
- They are free to use
- They are available for commercial use and can be built upon and customized
- They typically support community and academic use cases alike
- In most cases, they support an array of medical image file formats (including DICOM and NIfTI medical image file formats).
Now let’s take a closer look at some of the most popular open-source annotation tools on the market.
What are Some of the Most Popular Medical Imaging Open-Source Annotation Tools?
A wide range of open-source tools support, and were specifically created to, manage annotation projects for medical image datasets. In this article, we will focus on several of the most popular, including MITK Workbench, ITK-Snap, 3D Slicer, HOROS, OsiriX, and the OHIF viewer.
Here is a table outlining our top picks:
| Tool | Description | Key Features | Best For |
| MITK Workbench | Open-source software for medical image processing, annotation, and segmentation. | Cross-platform (Windows, Linux, macOS), GitHub source code, supports segmentation tasks, developed by German Cancer Research Center. | Researchers, PhD students, and teams working on segmentation-heavy medical imaging projects. |
| ITK-Snap | Focuses on medical image segmentation with manual and semi-automatic tools. | DICOM and NIfTI support, brush/paint tools, semi-automatic "Snake Interaction Mode," interpolation for segmentation. | Academic teams and practitioners starting with segmentation tasks. |
| 3D Slicer | Tool for visualization, processing, segmentation, and analysis of 2D and 3D medical images. | Extensive segmentation options (manual, semi-automatic), NIH-supported, multi-layered segmentation, 3D visualization. | Advanced imaging teams needing a robust tool for complex tasks, but prepared for a steeper learning curve. |
| HOROS | Medical imaging viewer and annotation tool optimized for macOS. | Cloud-based annotation, Apple-native 64-bit viewer, collaboration-friendly with report generation. | Mac users and teams needing an Apple-optimized tool for collaborative annotation projects. |
| OsiriX | Viewer and annotation tool with free (Lite) and paid (MD) options. | Lite version for basic use, MD version for FDA-cleared projects, 3D rendering, DICOM support. | Early-stage startups (Lite) or professional medical teams needing secure, FDA-compliant tools (MD). |
| OHIF Viewer | Cloud-based annotation tool developed by the Open Health Imaging Foundation. | DICOMWeb compliant, OpenID Connect standards, multi-modal image fusion, multiplanar reformatting, collaborative workflows. | Teams needing an advanced, cloud-based open-source tool with commercial-like features. |
MITK Workbench
MITK Workbench is a free open-source software for medical image processing, annotation, and segmentation. It’s based on The Medical Imaging Interaction Toolkit (MITK), “open-source software for the development of interactive medical image processing software.”
The source code is stored in GitHub and there is MTIK Workbench software that anyone can download and use for Windows, Linux, and Mac (macOS). Here’s more information about how you can use the MTIK Workbench for medical imaging annotation and segmentation projects.
MTIK, and the subsequent open-source workbench tool, were originally developed for and by PhD students and researchers in the Division of Medical and Biological Informatics (MBI) of the German Cancer Research Center.

ITK-Snap
ITK-Snap is another open-source medical imaging annotation tool – unlike some of the others we will cover in this article, it is focused exclusively on one step of the broader data annotation process: the segmentation task.
It was created as the result of a long-term collaboration between researchers at PICSL at the University of Pennsylvania, and the Scientific Computing and Imaging Institute (SCI) at the University of Utah, and as a result has a heavy academic following.
ITK-Snap’s main offering is manual segmentation tools (eg. brush and paint); it also provides a basic set of semi-automatic tools (mainly the ‘Snake Interaction Mode’) and complementary tools to the segmentation process (mainly the interpolation feature). It is a perfect fit and a very popular option for practitioner teams just starting out, and it supports DICOM and NIfTI medical image file formats.
3D Slicer
3D Slicer was created for the “visualization, processing, segmentation, registration, and analysis of medical, biomedical, and other 3D images and meshes.”
It comes with downloadable desktop software, can be used commercially, and enables access to an extensive development platform, and an active network of users. 3D Slicer helps medical imaging operations and data teams implement segmentation on multi-layered medical images, including 2D and 3D segmentation – tools available for segmentation include manual ones (eg. brush, drawing tool, and eraser) as well as a larger set of semi-automatic ones compared to ITK-Snap (eg. thresholding and level tracking).
3D Slicer also allows for basic tasks complementary to segmentation, including basic interpolation between slices, and filters. The main downside that teams often lament when using 3D Slicer for annotating images and files is the functionality set (which is often reported as being quite convoluted) and the steeper learning curve compared to other tools mentioned in this article.
For over 10 years, the US National Institutes of Health (NIH) has been a key contributor and supporter, and 3D Slicer has had over 1 million downloads since it was launched.

HOROS
HOROS is another free open-source medical imaging viewer and annotation software project. It is often a preferred tool when annotating with Apple computers and, not by coincidence, its stated goal is “to develop a fully functional, 64-bit medical image viewer for macOS”.
Annotation and ML teams can use the HOROS viewer and annotation tools to annotate medical images and videos, store them in the cloud, and create reports to document an annotation project collaboratively.
Several other medical and healthcare-related projects have contributed and are a part of HOROS, including OsiriX, OpenJPEG, OpenGL, VTK, ITK, DCMTK, GDCM, Grok, and Horos Cloud.
HOROS works with and is supported by technology partners in the healthcare sector as well as by donations from users.
OsiriX
Interconnected to, and a supporter of HOROS, OsiriX is another option that many teams opt for when looking for a labeling tool to get started with.
Initially fully open-sourced, OsiriX now offers either a ‘Lite’ version, which is available for free as a demo application, or OsiriX MD, which is a commercial version that you can use from $69.99 per month.
Similar to most open-source tools, OsiriX Lite is often leveraged by early-stage startups, proof-of-concept (POC) projects, and research work. Based on our many conversations with teams, a few key features are worth digging into when evaluating a tool like OsiriX Lite against others; specifically, its capabilities in regards to 3D rendering, as well as DICOM and collaboration features (which, depending on the use case, teams often cite as being limited).
On the other hand, one of the main benefits of OsiriX MD is to make up for the issue of security, which is one of the main challenges with open-source annotation tools (and which we’ll go through in a second more in-depth).
OsiriX MD is an FDA-cleared and CE II labeled tool, and this increased level of security and safety makes it a better option for teams annotating professionally (or undergoing or eyeing the FDA approval).

OHIF Viewer
The OHIF Viewer was developed by and is supported by the Open Health Imaging Foundation (OHIF), at the Massachusetts General Hospital (MGH), and is open-source software under an MIT license.
The OHIF Viewer is a tool for creating “custom workflows with user-friendly interfaces. Review cases and report results quickly, zero installation required.” It includes advanced visualization tools, and an easy-to-use annotation suite, and is compliant with DICOMWeb and OpenID Connect standards.
OHIF is an open-source annotation tool that comes close to commercial options, as it supports multi-modal image fusion, multiplanar reformatting, and more. It also comes with a cloud-based interface, making it easier to manage collaborative annotation projects.
Despite the various benefits of using open-source annotation tools for medical imaging projects, there are several downsides too.
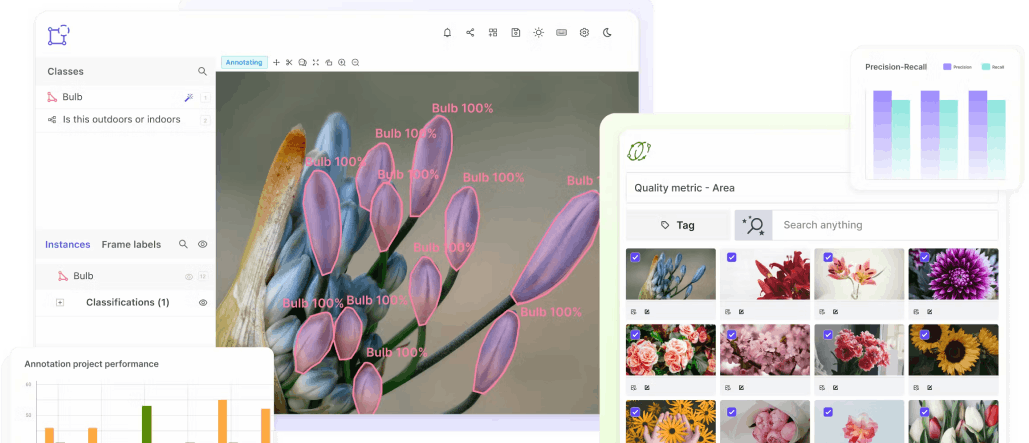
What are the Downsides of Using Open-Source Tools for DICOM Annotation?
As with any DICOM annotation tool, the ultimate goal of the labeling process is to provide high-quality data to the next step in the process; annotation is simply one stage, albeit a crucial one – in the case of building machine learning applications, once the datasets are labeled and annotated, you will be putting them to test by feeding them into a model (often a broader machine learning (ML) or computer vision (CV) model), then training and iterating on the training, until you are ready to launch a production-ready model and finally solve the critical objective you set out to achieve. Similarly, a hospitality software development company can apply such structured data processing and iterative enhancements to optimize guest experiences and operational efficiencies in travel and hospitality industries.
Open-source tools are often a great starting point when going from 0 to 1 with the annotation of medical imaging and video datasets, but, as we all know, they are inherently limited in their ability to achieve some of the more critical and powerful outcomes that teams face as they progress through the journey.
Throughout conversations with thousands of practitioners and leaders across the Medical AI community, we recurrently hear about numerous key downsides of open-source annotation software – these are often useful to consider ahead of time, in order for you to effectively plan out what your next steps in your data journey will be.
Below, we’ll dig deeper into the three main ones – scale, security, and collaboration – which you can also read more about in our blog here. These are:
- Scaling annotation activity with open-source tools is a big challenge. Open-source tools often come with a basic set of features (and hence are a perfect fit for many teams as they get started), but lack the wider set of needs that companies start to require at scale. For example, whereas it’s often a sensible setup for teams to collaborate on the annotation process over an open-source tool and back-and-forth emails, as the number of annotators and volume of data increases, in-app and real-time collaboration capabilities like tagging become key. Teams we work with often start to feel this pain point as they start to scale, and that’s when a more solid commercial option can help save resources, speed up the process, and also avoid errors and inaccuracy.
- Open-source tools inherently fall behind on security requirements. In most cases, open-source tools don’t benefit from the rigorous compliance standards of commercial tools, and by nature don’t include features like auditing, which more established companies require in order to achieve milestones like FDA approval. Many don’t have auditable data trails that can be monitored, tracked, and reported on, making it more difficult to achieve compliance with the FDA and HIPAA, or GDPR and CE certification in Europe.
- Free doesn’t always mean cost-effective. As the volume of annotations and annotators increases, the hidden costs of managing the process start to grow exponentially. At this stage, project leaders often find themselves needing to start being able to quantify, manage and measure their process and work; they need to be able to monitor and gain clear insight into the process and streamline operations on multiple spectrums. This is where limitations of open-source tools start to heavily affect their ability to achieve their objectives; two examples we hear often at this stage are frustrations around needing to write off a large percentage of time on non-value add tasks, as well as not being able to track how each annotator is performing, leading to poor process and output.
After getting started with open-source annotation software for their medical imaging dataset annotation projects, most teams tend to graduate to a commercial, proprietary tool that’s purpose-built to help take their project from 1 onwards. Tools like these are often easy to collaborate across, have best-practice security standards, and allow data operations and machine learning teams to scale their project cost-effectively (as well as help attain milestones like FDA clearance).

Encord is the leading annotation platform in medical AI, with a platform built to easily manage the annotation process, while allowing for the most complex of annotation tasks. At Encord, we have developed our medical imaging dataset annotation software in collaboration with data operations, machine learning, and AI leaders across the medical industry – this has enabled us to build a powerful, automated DICOM annotation suite, allowing for fully auditable data, and powerful labeling protocols.
A few of the successes achieved by the medical teams we work with:
- Stanford Medicine cut experiment duration time from 21 to 4 days while processing 3x the number of images in 1 platform rather than 3
- King’s College London achieved a 6.4x average increase in labeling efficiency for GI videos, automating 97% of the labels and allowing their annotators to spend time on value-add tasks
- Memorial Sloan Kettering Cancer Center built 1000, 100% auditable custom label configurations for its pulmonary thrombosis projects
AI-assisted labeling, model training & diagnostics, find & fix dataset errors and biases, all in one collaborative active learning platform, to get to production AI faster. Try Encord for Free Today.
Want to stay updated?
Follow us on Twitter and LinkedIn for more content on computer vision, training data, and active learning.
Frequently asked questions
DICOM annotation tools are software solutions designed to process and label medical images stored in the DICOM (Digital Imaging and Communications in Medicine) format. These tools help users classify, segment, and analyze medical imaging data, enabling its use in diagnostic workflows, research, and AI model development.
Open-source tools are a popular starting point for medical imaging annotation due to their cost-efficiency, customizability, and broad community support. They often support multiple file formats like DICOM and NIfTI, making them versatile for research and academic use.
Scalability Challenges: Managing large datasets and annotation teams is difficult.
Security Risks: They often lack compliance with FDA, HIPAA, or GDPR standards.
Hidden Costs: While free, managing processes at scale can introduce inefficiencies and indirect costs.
Project Scale: Can the tool handle increasing data volumes and annotators?
Compliance Needs: Does the tool support regulatory requirements like FDA or HIPAA compliance?
Collaboration Features: Are real-time and cloud-based collaboration supported?
Integration Potential: Can it be integrated into existing workflows?
Encord offers a user-friendly annotation platform designed to streamline the labeling process for medical imaging data. Our tools facilitate efficient collaboration among multiple centers, enabling consistent annotations across diverse datasets. This ensures that all researchers and healthcare professionals can easily access and utilize the labeled data for developing AI solutions.
When annotating challenging datasets like ultrasound loops or 3D MRI volumes, Encord encourages the use of standardized annotation methods to maintain clarity. For ultrasound, marking a region in one image can be propagated to others, while for MRI, annotating each slice with consistent techniques helps improve model understanding and reduces human error.
Encord aids in the evaluation of AI tools for medical imaging by providing a robust annotation platform that enables researchers to generate high-quality labeled datasets. This, combined with our expertise in the medical AI space, allows users to systematically assess the performance of AI solutions based on well-annotated images and clinical data.
Encord is equipped to manage various modalities of medical imaging, including X-rays, MRIs, and ultrasounds. The platform allows for specific annotation techniques tailored to each type, such as bounding boxes for fractures in X-rays or slice-by-slice annotations for 3D MRI data, ensuring accuracy and consistency in the training datasets.
Encord offers a scalable solution built specifically for high-demand environments, unlike many open-source platforms that may not meet the necessary performance and collaboration standards. Our platform is tailored to support the unique needs of companies in the medical field, enhancing both data quality and annotation efficiency.
Encord provides robust capabilities for managing and annotating hyperspectral imaging data, which is often challenging for many platforms. Unlike other solutions that may offer limited or cumbersome support, Encord is designed to handle hyperspectral data natively, ensuring a streamlined workflow for your specific needs.
While some teams may explore multimodal approaches by linking annotations with radiology reports, Encord primarily focuses on establishing ground truth through direct annotations. This ensures that the training process remains clear and aligned with the specific requirements of your project.
Encord specializes in providing advanced annotation tools specifically designed for medical imaging, including MRI and CT scans. Our platform streamlines the labeling process, ensuring high-quality annotations that are crucial for training machine learning models effectively.
Encord supports a variety of medical imaging data types including DICOM images such as X-ray, MRI, and CT scans. While the platform primarily focuses on these modalities, it is essential to clarify the specific requirements for each distinct application.
Absolutely! Encord is designed to manage annotations from different scans within the same study, allowing users to easily navigate and annotate across multiple images while maintaining context.
