The Comprehensive Guide to Medical Annotations
Medical annotations are more complicated than applying annotations and labels to non-medical images. In most cases, file sizes, formats, modalities, and the sheer volume of data is larger and more complicated than other image-based datasets. Medical annotations must be more accurate for training algorithmic models, patient outcomes, and healthcare plans.
Mistakes are costly in the medical profession. Patient healthcare plans, treatments, and outcomes depend on an accurate diagnosis. Medical image annotations are part of labeling anything from X-Rays to CT scans.
When applied to image or video-based datasets that are used to train computer vision, machine learning, and artificial intelligence (CV, ML, AI, etc.) models, medical annotations are integral to new treatment innovations across the healthcare sector.
In this article, we provide more detail about the medical image annotation process, including healthcare use cases, best practice guidelines, and considerations medical ML teams need to factor in when annotating images and videos.
What are Medical Image Annotations?
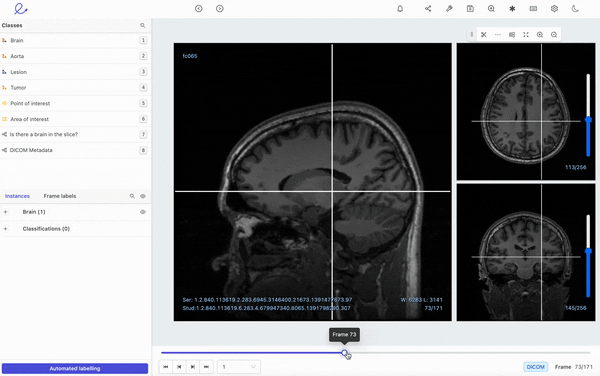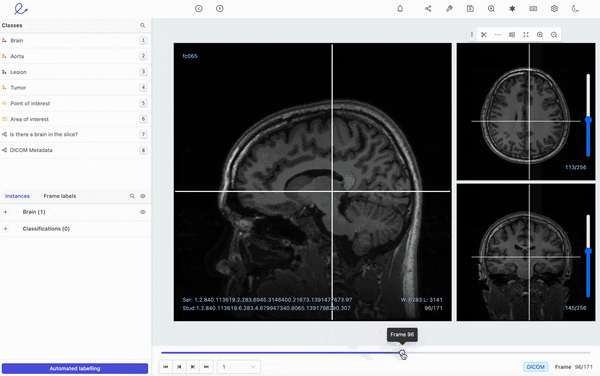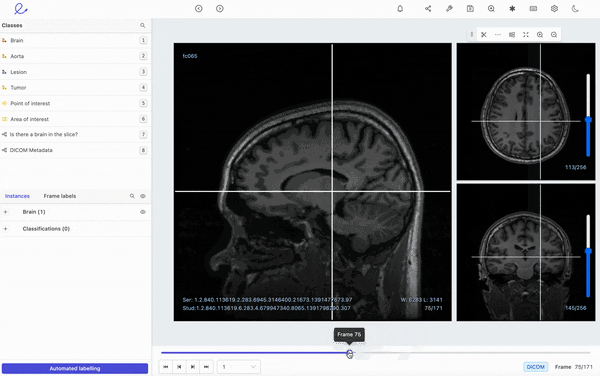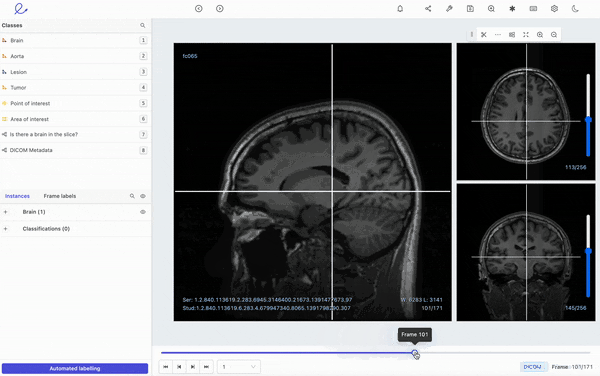
Encord's DICOM annotation tool
Computer vision models and other algorithmic models, such as artificial intelligence (AI), and machine learning (ML), rely on accurately annotated and labeled datasets to train them. In a medical environment, the annotations and labels need to be even more precise to produce accurate outcomes, such as diagnosing patients.
Accurate annotated examples of medical images are crucial for training and making a model production-ready. Annotations usually are initially provided by experts in the relevant medical specialism. Then annotation teams and AI-powered tools normally take over to annotate vast datasets based on the labels created.
The best example of this is a radiologist who uses an annotation platform to note down their opinion of a scan, which in turn trains the neural network accordingly. Companies can build their own labeling platform or take advantage of third-party medical imaging labeling tools (take a look at this blog for the pros and cons of each approach). Whatever you decide to do, the better your approach to labeling your DICOM or NIfTI images, the better your model will perform.
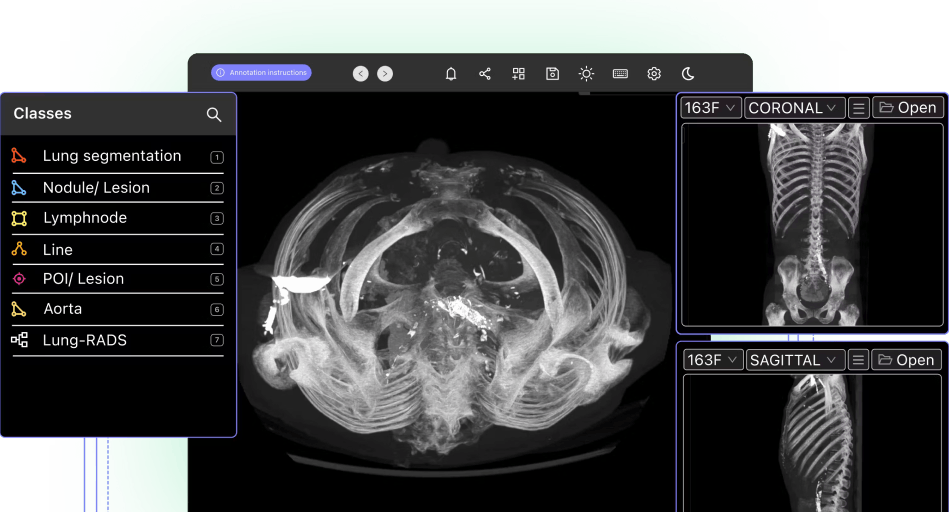
How does Medical Image Annotation Compare With Regular Image Annotation?
Medical image annotation is more complex than annotating datasets filled with (non-medical) images, such as JPGs or PNG files.
Medical data ops and annotation teams have much more to consider, such as regulatory compliance, layered file types, 2D, 3D, and even 4D formats, windowing control settings, and much more.
Here’s a table to explain some of the challenges of medical image annotation:
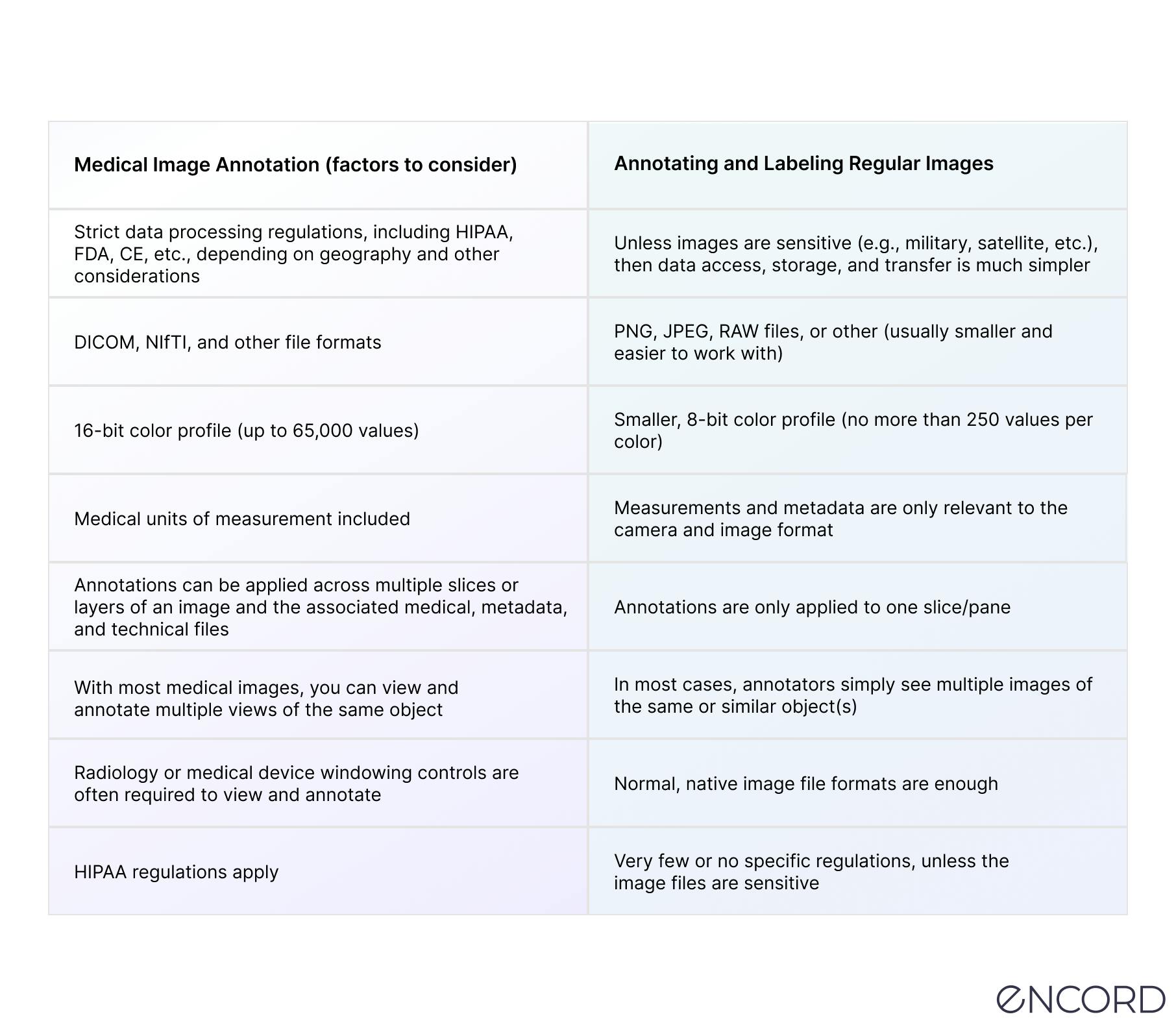
Now let’s take a look at medical annotation use cases and the numerous ways annotations and labels can be applied for computer vision models in the healthcare sector.
Medical Image Annotation Use Cases
There are hundreds of use cases for medical annotations and labeling across dozens of specialisms and healthcare practices, including the following:
Pathology
For the vast majority of diseases, most of the diagnostic capabilities come from various scans and images that are taken by highly specialized medical equipment. By labeling these scans accurately, we can train machine learning models to pick up those diseases themselves, reducing the need for human involvement.
Radiology
Radiology is one of the most common use cases for medical image annotations. Mainly because of the vast number of images this medical field generates, with dozens of radiology modalities, including X-ray, mammography, CT, PET, and MRI.
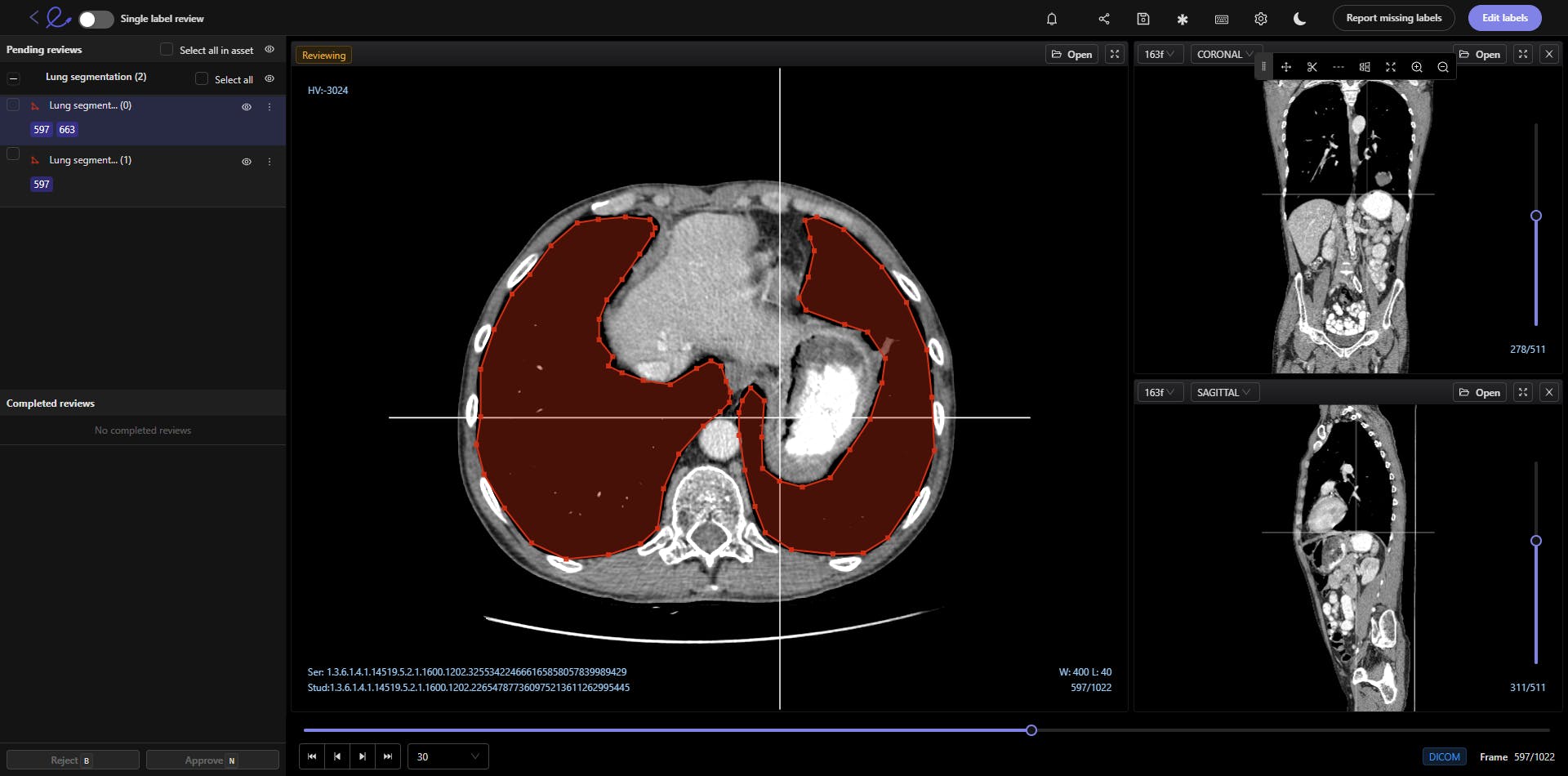
With the right annotation tool, medical data ops and annotation teams can benefit from a PACS-style interface to make native DICOM and NIfTI image rendering possible. Plus, annotation tools should come with customizable hotkeys and other features for multiplanar reconstruction (MPR) and maximum intensity projection (MIP).
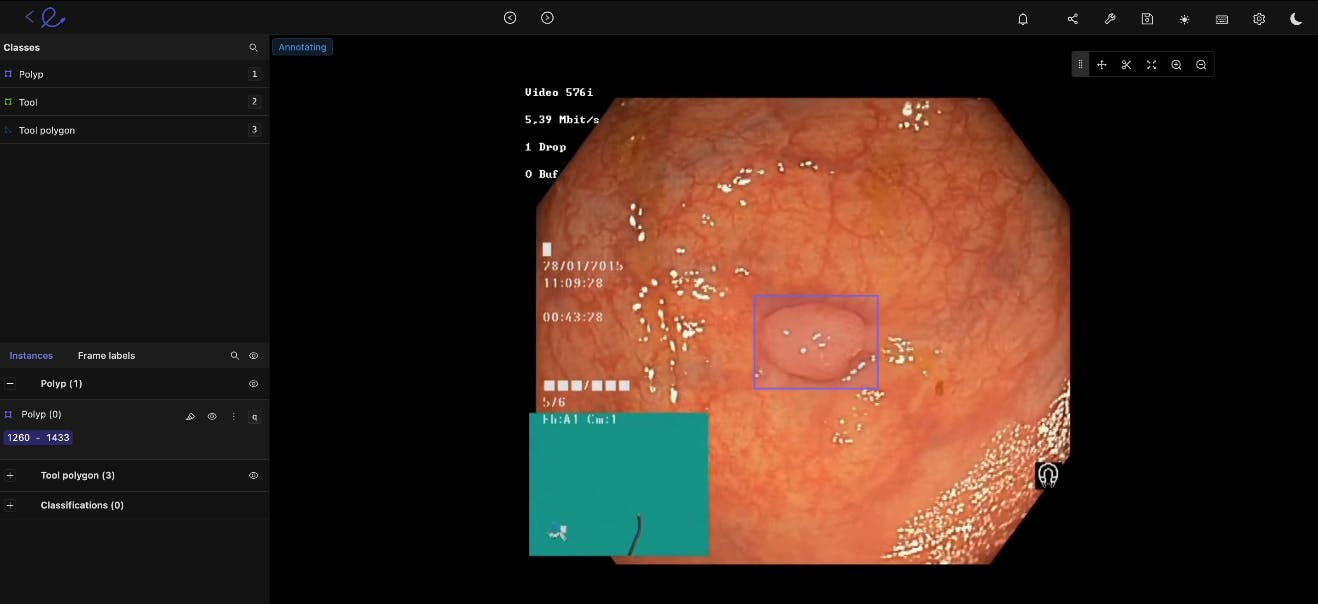
Gastroenterology
You can improve video yields and accelerate GI model development with the right annotation tool. For gastroenterology model development, you need an annotation tool to support native video uploads of any size or length. This proves especially useful for annotation and computer vision work designed to detect cancerous polyps, ulcers, IBS, and other conditions.
Histology
Medical image annotation is equally useful for histology, giving annotators and medical ops teams the ability and tools to Increase micro and macroscopic data labeling protocols and training datasets.
With the right annotation tools, you should have inbuilt support for the most common and widely-used staining protocols (including hematoxylin and eosin stain (H&E), KI67, and HER2). Plus, it’s time and cost-saving to use an annotation platform compared to booking out expensive microscopy workstation time.
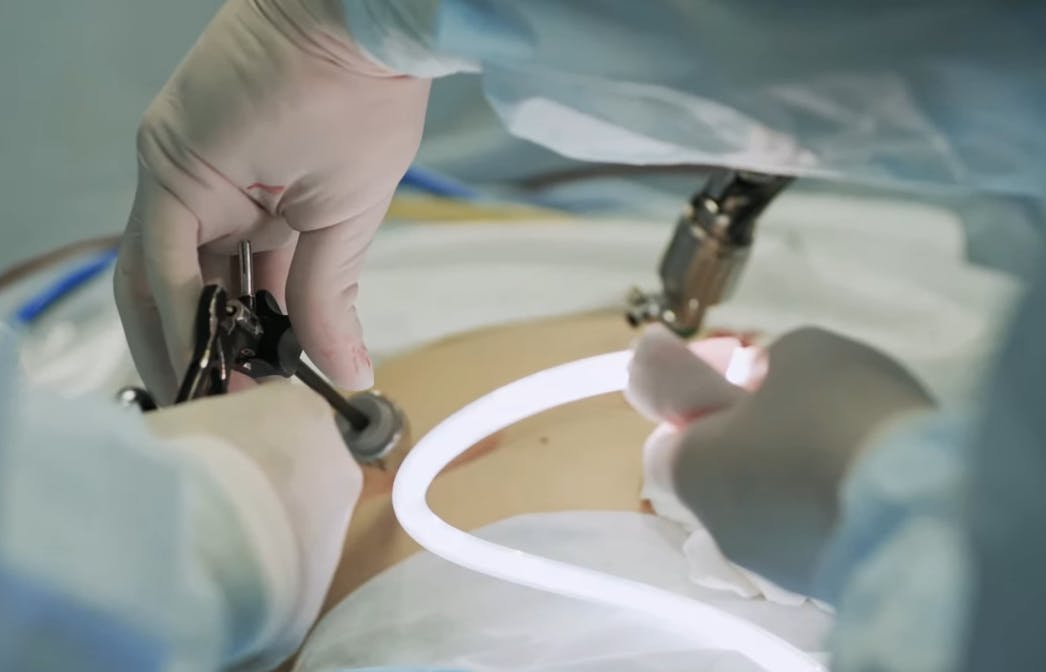
Surgical
Surgical AI models benefit from faster, AI-powered automated annotations. For this, medical data operations teams need a medical-grade video annotation and clinical operations platform designed for surgical intelligence use cases.
Cancer Detection
Cancers are notoriously challenging to diagnose, so medical image annotation can help us train models that spot them earlier and more accurately than humans can – which can make a huge difference to patient outcomes. When computer vision models are used to screen for the most common cancers automatically, we can drastically improve early detection and treatment plans and outcomes.
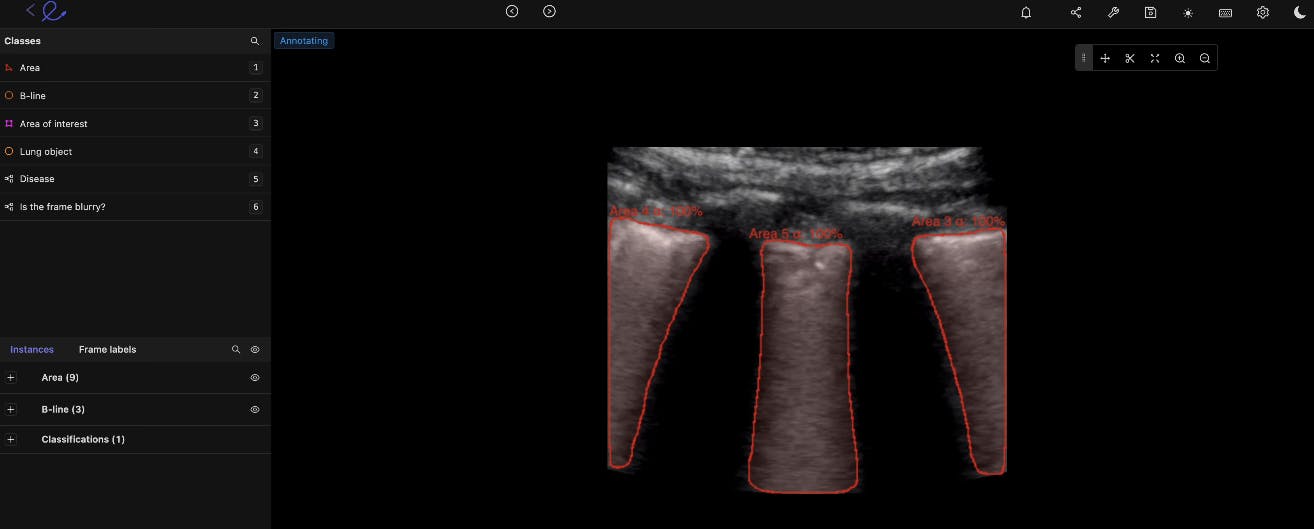
Ultrasound
By annotating ultrasound images, we can use artificial intelligence (AI) to pick up higher levels of granularity for things like gallbladder stones, fetal deformation, and other diagnostic insights. The quicker we understand what we’re dealing with, the better the care will be.
Microscopy
In medical research, we rely a lot on what we can examine under a microscope to understand what’s happening at the lowest level of abstraction. By labeling these images and applying them as a training dataset, we can push medical research forward and scale our impact as a result.

The beauty of machine learning is that there are many more use cases to discover as we start to work with this data and let the algorithms do their thing. This is at the forefront of the future of medicine, and the quality of the annotations will be a significant factor in how things evolve.
 Read more about the top 6 AI trends in healthcare.
Read more about the top 6 AI trends in healthcare. 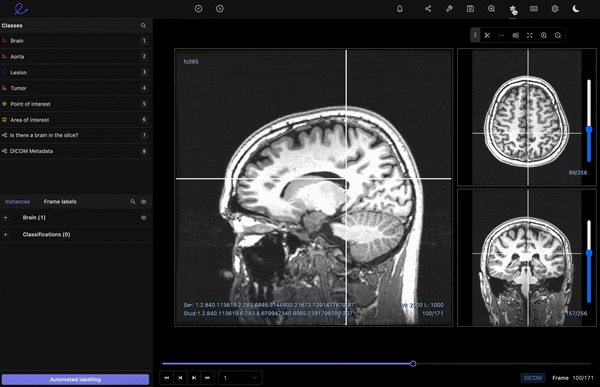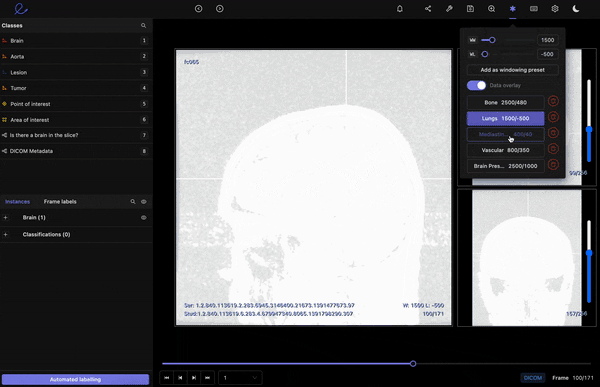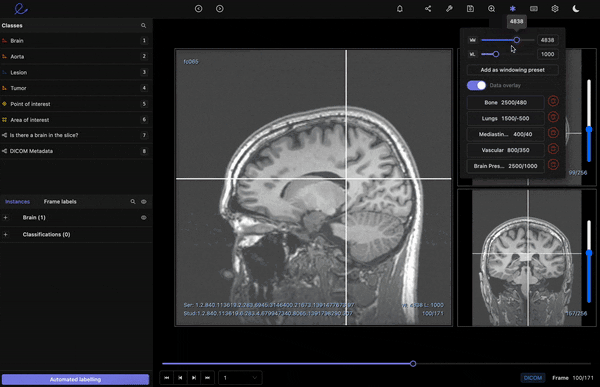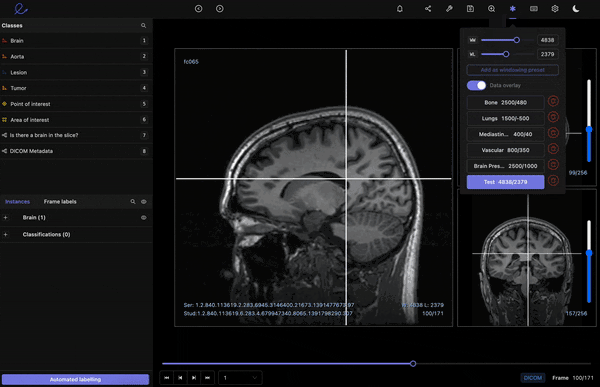
Windowing preset feature on the Encord medical image annotation tool
Essential Considerations When Preparing Healthcare Imaging Data for Computer Vision
Medical image annotation relies on high levels of precision because of the complexity involved and the stakes under which these models will be used. In order to pass FDA approval and make it to production, the data must be of the highest quality possible – to adhere to regulatory guidelines and create a stronger and more effective machine learning model.
To do this at scale, organizations need to make it as easy and intuitive as possible for annotators to capture the required information. The time of these experts is costly, so the more efficient and frictionless the annotation process, the better quality data you’ll get and the more you can control costs.
Four key considerations should be prioritized when you’re tasked with annotating medical imaging data, or managing an annotation team:
Dataset volumes, sizes, and file types
As with any machine learning project, the more data you have to train with, the better the model will perform. This assumes a certain level of data quality, of course, but wherever you can – you should try to increase the size of the training set as much as possible.
Data Distribution & Diversity
In the medical field, there is tremendous diversity in terms of human bodies, and that needs to be reflected in your data. This diversity is crucial if you want your model to be effective in the real world. It would be best to ensure sufficient distribution across demographic factors like age, gender, geography, hospitals, previously diagnosed conditions, and other relevant details.
 Read more: Encord’s guide to medical imaging experiments and best practices for machine learning and computer vision.
Read more: Encord’s guide to medical imaging experiments and best practices for machine learning and computer vision. Data Formats
Medical images can come in various formats, including DICOM and NIfTI images, CT (Computed Tomography) scans, X-Rays, and Magnetic Resonance Imaging (MRI) files.
Your annotation process needs to handle these formats natively so that you don’t lose any detail or information along the way. This ensures that you’re getting the most out of your workflow and know it can fit into the existing medical system in which you want to innovate.
Data Visualization
When images are delivered in 3D, you need to ensure you view the files in the same format to get the full picture of whatever medical image is annotated. You want to provide the annotator with everything they need to provide an accurate evaluation, which means that you need to be thoughtful and intentional about how you present the images to them.
It’s equally important to consider the different views and volumes of medical images, such as 2D and 3D. In most cases, you need both to assess what’s happening in the images accurately.
Image types such as MRI, CT, and OCT scans can be viewed in a number of ways, such as using the sagittal, axial, or coronal planes. However, for ML purposes, it’s more time and cost-effective to pick and annotate those to train a model.
This list is not exhaustive, but it should give you a sense of what to consider when building a medical image annotation workflow.

Brush selection tool for annotating DICOM and NIfTI images in Encord
How to Implement Medical Image Annotations?
Here are the steps you need to take to implement medical image annotations for a computer vision project.
Sourcing Data
Medical image or video datasets can be sourced in numerous ways. There are dozens of open-source medical datasets. Healthcare organizations might have their own in-house data sources they can tap into and access. Or you will have to buy datasets from hospitals and healthcare providers.
It depends on how much data you need (images, videos, etc.), and the health issues you’re investigating and training an algorithmic model on.
Budgets, timescales, and the resources you invest in annotating and labeling datasets also play a role. As do the tools you’re going to use, as automated tools accelerate the medical image annotation process.
 Need to source medical imaging data quickly and for free? Here are the top 10 free healthcare datasets.
Need to source medical imaging data quickly and for free? Here are the top 10 free healthcare datasets. Preparing Medical Image Datasets
Once you’ve sourced these datasets, the images and videos within need to be cleaned and prepared for the annotation process.
If your AI model is being trained as part of a commercial project aiming for FDA approval, then it’s essential that patient identifiers are removed from tags and metadata. You need to split and partition the dataset, ideally keeping it in a separate physical location to ensure FDA approval is easier to achieve.
Create the Annotations and Labels
Next, the annotations and labels need to be created. In most cases, when a project is for a medical use case, healthcare professionals should create the labels. Especially if the annotation and labeling work is being outsourced to non-experts; otherwise, you risk the entire project by hoping annotators know what they’re looking at.
Once the labels are ready, and ideally pre-populated in a small sample of the dataset, then the annotation work can begin. Whether in-house or outsourced, starting this with smaller selections from the overall dataset makes sense. This way, especially initially, annotator outputs can be more effectively measured against data labeling key performance indicators (KPIs).
A quality assurance loop can be cycled through a few iterations until enough annotations and labels have been applied to validate and test a dataset sample, to start training the model.
Validate and Test A Dataset Sample
Medical imaging computer vision model training involves a continuous series of experiments. How much data scientists are involved depends on the way a model is being trained (e.g., unsupervised, semi-supervised, self-supervised, human-in-the-loop, etc.), and the outcomes of the validation and test dataset samples.
In some cases, you might need to re-do the labels and annotations until they’re more accurate. Or very least, a percentage of them might need to be improved and enhanced until the dataset is ready to be fed into the ML model.
Data Security & Regulatory Compliance (HIPPA, FDA, CE) for Medical Image Annotation Tools
Alongside all the considerations about the quality of the data being captured and the efficiency of the process, we also need to think carefully about the security of the images that you’re annotating. The labeling tool that you’re using needs to conform to the most up-to-date security best practices to reassure your stakeholders and customers that you’re taking medical data security seriously.
There are two key regulatory frameworks that you should be aware of here if you’re looking for an external medical image annotation tool:
- SOC 2 defines criteria and benchmarks for managing customer data and is measured through an external audit that evaluates data security practices across the board.
- HIPAA (the Health Insurance Portability and Accountability Act) is a U.S. Federal law that protects sensitive patient health information. This is a non-negotiable for whoever is providing your data labeling tool.
Depending on why an algorithmic model is being developed, you may also need to consider US Food & Drug Administration (FDA) compliance. We’ve got a guide for getting AI models through the FDA approval process.
For AI companies or healthcare organizations operating in Europe, there are also GDPR and European Union (EU) CE regulations to think about. Although they operate in similar ways to US data protection and healthcare-specific laws, it’s important to ensure data handling, processing, security, and audit trails are compliant with the relevant legislation.
As well as the data security credentials of your annotation tool provider, you also need to control the permissions available to your annotators carefully. You want to have very granular access controls so that they only see the absolute minimum that they need in order to do their job.
All of this should be tied up in a product where you retain the rights to your data and models – which simultaneously protects your IP and makes it easier to ensure high-quality data protection from the source all the way to final outputs.

Different Types of Tools for Medical Image Annotations & Labels
Medical imaging, healthcare, and annotation teams have three main options for selecting medical annotation tools:
- Open-source annotation tools;
- In-house annotation tools,
- Or powerful third-party annotation tools and platforms.
The first two, open-source and in-house, have several limitations, especially when it comes to data compliance, scalability, collaborative workflows, and in some cases, audit trails or the lack thereof.
In-house tools are also notoriously expensive and time-consuming to build and maintain.
On the other hand, with a powerful third-party annotation tool, you can be up and running quickly, have complete data auditability, and compliance, and benefit from powerful medical annotation and collaborative workflow features. Including quality control and quality assurance.
 Here are three ECG annotation tools you can use.
Here are three ECG annotation tools you can use. Medical Image Annotations With Encord
At Encord, we have developed our medical imaging dataset annotation software in collaboration with data operations, machine learning, and AI leaders across the medical industry – this has enabled us to build a powerful, automated image annotation suite, allowing for fully auditable data, and powerful labeling protocols.
Encord comes complete with Encord Active, Encord’s Annotator training module, and an extensive range of features specifically for medical image annotations, labeling, AI-assisted labeling, quality assurance, and workflows.
Case studies from projects that Encord has been deployed in for medical annotations include the following:
- Floy, an AI company that helps radiologists detect critical incidental findings in medical images, reduced CT & MRI Annotation time with AI-assisted labeling
- RapidAI reduced MRI and CT Annotation time by 70% using Encord for AI-assisted labeling.
- Stanford Medicine cut experiment duration time from 21 to 4 days while processing 3x the number of images in 1 platform rather than 3
Ready to automate and improve the quality of your data labeling?
Sign-up for an Encord Free Trial: The Active Learning Platform for Computer Vision, used by the world’s leading computer vision teams.
AI-assisted labeling, model training & diagnostics, find & fix dataset errors and biases, all in one collaborative active learning platform, to get to production AI faster. Try Encord for Free Today.
Want to stay updated?
Follow us on Twitter and LinkedIn for more content on computer vision, training data, and active learning.
Join our Discord channel to chat and connect.
Frequently asked questions
Encord provides tailored solutions that cater to the specific needs of pediatric applications in medical imaging. The platform's annotation capabilities are designed to accommodate the unique challenges posed by pediatric images, helping healthcare professionals to annotate accurately and efficiently.
Encord prioritizes security and user experience in its design. The platform is built to be user-friendly, ensuring that radiologists and other professionals can easily navigate it. Additionally, we adhere to strict security standards to protect sensitive medical data throughout the annotation process.
Encord offers a robust annotation platform that simplifies the labeling of customer images. It enables teams to handle large volumes of data efficiently, ensuring that the annotation process is accurate and meets customer needs, especially when customers struggle with providing properly labeled data.
Encord streamlines the annotation process by offering tools that not only focus on the drawing of boxes and lines but also improve the organization of tasks. This allows for efficient workflows, making it easier to manage larger studies without overwhelming the medical professionals involved.
Encord provides tools specifically designed for NLP projects, enabling users to label and annotate text data efficiently. Our platform supports complex analyses, such as identifying medical conditions from health records, which is crucial for improving patient outcomes.
Yes, Encord is well-suited for repetitive labeling tasks, making it ideal for projects like medical device development. The platform allows users to efficiently manage consistent image types and tasks, helping teams maintain productivity with minimal complexity.
Encord offers frameworks for active learning that aid in selecting high-quality images for annotation. By utilizing models to identify difficult cases and screen out poor-quality images, Encord ensures that only the most relevant data is sent for labeling.
Encord provides advanced annotation tools, including features like AutoSAM and Magic Select, which facilitate efficient segmentation of images. These tools are designed to streamline the annotation process, making it more intuitive and user-friendly.
The label editor in Encord offers a user-friendly interface for annotators, allowing them to easily create and manage labels for different imaging modalities. It includes tools for precise annotations, ensuring high-quality data for AI training.
Encord provides capabilities to annotate CBCT DICOM files in 3D, enabling users to efficiently label complex medical images. This feature is crucial for teams looking to enhance their annotation strategies in the medical domain.
